Aaron N. Waite and Dr. Jack W. Sites Jr., Zoology
Introduction
The genus Gymnodactylus is a group of small lizards belonging to the family Gekkonidae, which occur in the east and central areas of Brazil. The species G. darwinii is restricted to the Atlantic Forest and ranges from the northeastern state of Bahia south 1500 miles along the coast to the northern region of the southeastern state of São Paulo.
Different populations of similar organisms can be recognized as distinct species on the basis of many criteria including morphology, chromosomes, and genetic divergence of Mendelian and cytoplasmic loci (1). These differences are supported by an ecological distinction between the species, whether it is a large geographical separation or a difference of environmental niche that impedes gene flow between the two. The genetic and morphological divergence can be used as a standard to reference differences between geographically discrete populations within a species.
The purpose of this study is to investigate the genetic divergence of mitochondrial DNA sequences between three separate populations of G. darwinii, collected along its range in the Atlantic forest of Brazil, in order to test the hypothesis that all populations are conspecific.
Material and Methods
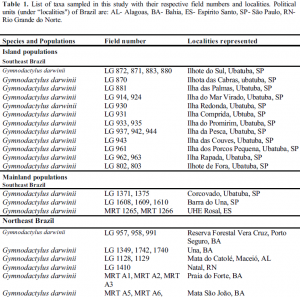
Forty-two specimens were used to reconstruct a molecular phylogeny of Gymnodactylus darwinii. The sampling includes three geographically distinct populations across a wide range of the Atlantic Forest of Brazil. The first is found in the northern most range in the states of Bahia, Alagoas, and Rio Grande do Norte, and the second is a mainland region in the states of São Paulo and Espírito Santo, in southeast Brazil. The third is composed of specimens from 12 small land-bridge islands off the coast of São Paulo (Table 1). The gekkos Phyllopezus pollicaris and Hemidactylus mabouia were used as outgroups.
Total genomic DNA was extracted from liver and tail samples according to Fetzner (2). About 700bp from the mitochondrial gene cytochrome b were amplified using universal primers (3) by the polymerase chain reaction (PCR). Sequences were fractionated on an ABI-377 Automated Sequencer in the DNA Sequencing Center on the sixth floor of the Widtsoe Building, and edited and aligned using the computer program Sequencher 3.1.1(Gene Codes Corp., Inc., 1995). Phylogenetic analysis under the optimality criteria of minimum evolution was performed in PAUP* version 4.0b4a (4). The GTR+Γ+I was selected as the best fit model of evolution for the cyt b sequences (5), and then used to reconstruct the phylogeny for G. darwinii. Confidence in the recovered nodes in the topology was assessed by non-parametric bootstrap (6).
Results and Discussion
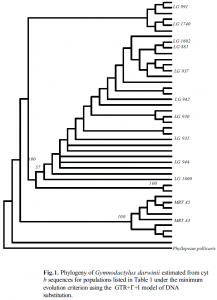
Figure 1 presents the phylogeny estimated from the analysis of cyt b sequences collected from different populations of G. darwinii from the mainland and islands in the Atlantic Forest in Brazil. Four major clades were recovered. The first is a large clade including all the specimens from the 12 small land-bridge islands off the coast of São Paulo, and the adjacent mainland populations. The second is a small clade nesting the population from the mainland in the state of Espírito Santo as the sister group of the São Paulo clade. Two other well supported clades (100% bootstrap) nested the populations from the northeast of Brazil. These results indicate a lack of conspicuous population structure for G. darwinii from the islands and mainland in the southern part of its range (São Paulo). This can be due to the relatively short period of time (11,000 years) since these islands were isolated from the mainland and each other (7). Nevertheless, the populations from the northern part of the range of G. darwinii are completely separated and split into two distinct clades despite the close geographic distribution. This corroborates with the chromosomal differentiation (8) already found between some populations from the islands mainland complex in São Paulo (2n=38) and those from mainland regions in the northeast of Brazil (2n=40). This study suggests that the populations from the northeast might represent distinct evolutionary units.
References
- Sites, J.W.; Crandall, K.A. (1997). Testing species boundaries in biodiversity studies. Conservation Biology 11: 1289-12971.
- Fetzner, J. (1999). Extracting high-quality DNA from shed reptile skins: a simplified method. Bio Techniques 26: 1052-1054.
- Palumbi, S. R. (1996). Nucleic acids I: the polymerase chain reaction. Pages: 205-247 in Molecular systematics, 2nd Edition (D. M. Hillis, C. Moritz, and B. K. Mable, eds.). Sinauer Associates, Inc., Sunderland, Massachusetts
- Swofford, D. L. (1998). PAUP*: Phylogenetic analysis using parsimony (*and other methods), Beta Version 4.0.b4a. Sunderland, Mass: Sinauer.
- Posada, D.; CrandalL, K.A. (1998). Model test: testing the model of DNA substitution. Bioinformatics 14: 817-818.
- Felseinstein, J. (1985). Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39: 783-791.
- Vanzolini, P.E. (1973). Distribution and differentiation of animals along the coast and in the continental islands of the state of S. Paulo, Brazil. I. Introduction to the area and problems. Papeis Avulsos, Zool., São Paulo 26:281-294.
- Pellegrino, K.C.M. (1998). Diversidade cariotípica e evolução cromossômica em lagartos das famílias Gymnophthalmidae e Gekkonidae (Squamata): evidências baseadas em coloração diferencial e hibridação in situ fluorescente (FISH). Genetics and Molecular Biology 21(3): 418-419 (Thesis abstract).
- Faculty mentor, Brazil: Dr. Katia C. Pellegrino, Departamento de Biologia Universidade de São Paulo, SP, Brazil