Steven Alan Smith and Dr. Laura Bridgewater, Zoology
Osteoarthritis is one of most prominent illnesses that plague our world today. It involves the degeneration of cartilage between joints due to a reduction of collagen fibrils. The type XI collagen gene (Col11a2) is one of a few genes responsible for the production of the collagen fibrils. It is expressed wherever cartilage is present. It is expressed in these areas because of specific expression factors (proteins) that enhance the production of collagen. Once these expression factors are known, a possible genetic remedy can be manufactured to aide in curing osteoarthritis or other skeletal abnormalities.
Previously, Dr. Laura Bridgewater discovered an enhancer region in the first intron of the Col11a2 gene. An enhancer region is a sequence of DNA to which proteins bind and enhance the expression of certain genes. Transfection assays assisted in making this discovery. For this process a circular piece of DNA (plasmid) is manufactured containing the potential enhancer site. It also contains a reporter gene that if expressed, will produce enzymatic activity that can be detected by a machine called the luminometer. Therefore, the expression of that gene and hence the enzymatic activity is dependant on whether the potential enhancer region does indeed contain an enhancer sequence. Dr. Bridgewater concluded that the potential region did include an enhancer site known as the FG region (1).
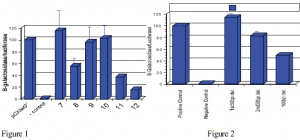
Further tests showed that the FG enhancer was possibly accompanied by other enhancer sequences in close proximity to the FG site. Our work was to identify the unknown potential enhancer regions near the FG site. We deleted six 100bp (base pair) regions of the DNA and observed the enzyme activity of the plasmid excluding one deleted region at a time. If a deleted region contained an enhancer site then enzyme activity would be low. If it did not, then enzyme activity would be high. The eleventh region showed low activity because it housed the FG enhancer site (Figure 1). Two other regions (numbers 8 and 12) were also low in activity indicating that either one or both may house the unknown enhancer sites. Region 12 has since been ruled out as containing an enhancer site and my research has involved region 8.
To verify that region 8 contained an enhancer site for the Col11a2 gene, the entire 100 base pairs of site 8 were cut in half and similar transfection assays were performed with each half. This was done to determine if the enhancer region was located in the first 50 base pairs or in the second. After much delay due to contamination, results were finally obtained (Figure 2.) These results were discouraging at first because enhancer activity of any of the 50 base pair deletions did not drop completely or to the same level as when the entire 100bp were deleted out. This made it difficult to determine where the enhancer region was located, if it existed at all. However, the results were interesting in the following way.
First of all, when the first 50bp of region 8 are deleted enzyme activity increases. However, when the second 50bp are deleted the activity decreases below the positive control but not as far below as the entire 100bp. These results are similar to data produced by another group of researchers who identified both an enhancer sequence and an inhibitor sequence near a collagen gene (2). An inhibitor sequence does exactly the opposite as an enhancer by decreasing expression of a gene. Hence, if an inhibitor is deleted from the DNA sequence then activity will increase, which is the case for the first 50bp of region 8. In addition, the DNA sequence of the inhibitor identified by the group is also found in the first 50bp of region 8.
Secondly, this group also identified another sequence that enhanced activity near the sequence that inhibited activity. This is also consistent with my data because when the second 50bp is deleted activity decreases. This data leaves me with two questions. First, if the second 50bp does contain the enhancer site, then why does activity decrease more when the entire 100bp is deleted than when the second 50bp is deleted? Secondly, if the first 50bp contain an inhibitor then why would activity decrease at all when the entire 100bp is deleted?
I hypothesize that there is indeed an enhancer site but it is located overlapping the two 50bp deletions. This is why activity decreases much more when the 100bp are deleted compared to when the second 50bp are deleted. I also hypothesize that there is indeed an inhibitor region in the first 50bp but that it is inactive when the enhancer site is active. My plans are to continue this research and find answers to these questions.
References
- Bridgewater, L.C., V•ronique Lefebvre, and Benoit de Crombrugghe. 1998. Chondrocytespecific Enhancer Elements in the Col11á2 Gene Resemble the Col2á1 Tissure-specific Enhancer. J Biol Chem, Vol. 273, Issue 24, 14998-15006.
- Murray, Darryl, Patricia Precht, Richard Balakir and Walter E. Horton. 2000. The Transcription Factor äEF1 Is Inversely Expressed with Type II Collagen mRNA and Can Repress Col2á1 Promoter Activity in Transfected Chondrocytes. J Biol. Chem, Vol. 275, No. 5, 3610-3616.
- Bridgewater, L.C., V•ronique Lefebvre, and Benoit de Crombrugghe. 1998. Chondrocytespecific Enhancer Elements in the Col11á2 Gene Resemble the Col2á1 Tissure-specific Enhancer. J Biol Chem, Vol. 273, Issue 24, 14998-15006.
- Murray, Darryl, Patricia Precht, Richard Balakir and Walter E. Horton. 2000. The Transcription Factor äEF1 Is Inversely Expressed with Type II Collagen mRNA and Can Repress Col2á1 Promoter Activity in Transfected Chondrocytes. J Biol. Chem, Vol. 275, No. 5, 3610-3616.