Becky A. Shiozawa and Professor R. Paul Evans, Zoology
Current mitochondrial DNA techniques using restriction fragment length polymorphisms (RFLPs) are still unsuccessful in determining phylogenetic relationships between subspecies of cutthroat trout (Oncorhynchus clarki), such as the relationships between Snake River finespot cutthroat trout and Yellowstone cutthroat trout, Bear River Bonneville cutthroat trout and Yellowstone cutthroat trout, and Lahontan cutthroat trout and its subspecies. Due to these groups distinct geological or ecological separation and the relatively rapid rate of mtDNA evolution, it is likely distinctive markers exist for these subspecies within mtDNA. Identification of such markers would be valuable for identification and future research on the subspecies.
A phylogenetic chart using mtDNA sequencing data for differentiating between the various cutthroat trout species was generated. This chart was compared to a phylogenetic tree generated using other techniques for correlation. For this study, emphasis was placed on the Lahontan cutthroat trout, the finespot cutthroat trout from the Snake River, and the Bonneville cutthroat trout from the Bear River.
Samples were obtained from the Monte L. Bean Life Science Museum archives of fish tissues and DNAs. Three populations from each subspecies of cutthroat trout with approximately five fish per population were used (total of 95 samples). DNA was isolated from tissue samples using techniques established in Dr. R. Paul Evans’ laboratory. There were some difficulties in isolations due to the poor quality of the tissue samples from improper preservation or age of the tissue. The cytochrome B and ND1 regions of the mtDNA genome were amplified using polymerase chain reaction (PCR). There were difficulties in PCR amplification for many of the samples. Different DNA templates were tried and as well as lowering the annealing time programmed in the PCR machine. Temperatures tried include 550C, 500C, and 470C. As to this date, there are still several populations that have failed to amplify. Further work on these populations will be continued and will not be contained in this report. Finished PCR products were gene cleaned, sequenced, run through a sephadex column, and submitted to the BYU DNA sequencing center for analysis. The finished product was the broken down genetic code for the mtDNA regions (see figure 1). Using Sequencher 3.1, the samples’ genetic code was aligned to match up as accurately as possible and was checked manually for errors or corrections. The finished code was run through the computer program PAUP 4.0 to generate a phylogenetic tree for the CYTB and ND1 regions.
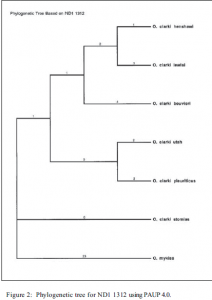
The trees generated do correlate with data generated from nuclear markers and indicate the use of CYTB and ND1 mtDNA sequencing to identify subspecies of cutthroat trout. The phylogenetic tree generated from CYTB data indicates its use in differentiating O. clarki clarki from either O. clarki bouvieri or O. clarki utah. However, it is insufficient in differentiating between O. clarki bouvieri and O. clarki utah. Data from the ND1 region generated a more diversified phylogenetic tree (see figure 2). Using this region, O. clarki stomias is distinct from the other groups. O. clarki bouvieri is also differentiable. Further work will need to be done in order to differentiate between O. clarki henshawi and O. clarki lewisi, as well as between O. clarki utah and O. clarki pleuriticus.
This study demonstrates the usefulness of mtDNA in the identification of some subspecies of O. clarki although further research is still necessary for total differentiation. Other regions of the mtDNA genome may be examined for further differentiation between the subspecies. However, these results do allow for further identification within this group and advances research in this area.
Literature Cited
- Shiozawa, B. A., unpublished data.
- Shiozawa, D. K. and R. N. Williams. 1992. Geographic variation, isolation and evolution of cutthroat trout (Salmo clarki). Final Report to the National Geographic Society. Grant #3653-87.