James A. Robertson and Dr. Michael F. Whiting, Zoology
The Cucujoidea, a very diverse beetle superfamily, includes mycophagous and wood consuming beetles. Within the Cucujoidea is the family Erotylidae (pleasing fungus beetles). The Erotylidae includes 125 genera and approximately 2,500 species, and has a worldwide distribution. Erotylids are known for their striking coloration, including black with yellow, red, orange and even pink and purple (Boyle 1956). The current hypothesis of erotylid phylogeny is based exclusively on adult morphology and was not formulated using rigorous phylogenetic methodology. Hence phylogenetic relationships of the major lineages within this family are dubious. Since erotylid beetles exhibit such striking aposematic coloration, a well-supported phylogeny, constructed independent from their morphology, is a critical step towards understanding the evolution of this coloration and their association with fungi. This research is a first step in the exploration of the utility of molecular data for identifying major lineages within the Erotylidae.
Ingroup taxa consist of eight erotylid exemplars, representing three of the five subfamilies. Outgroup taxa were selected from the superfamilies Cucujoidea, Tenebrionoidea, Bostrichoidea, and the basal beetle lineage Caraboidea. DNA was extracted from thoracic or leg muscle tissue using standard phenol/chloroform extraction protocols. 18S rDNA was amplified via the polymerase chain reaction (PCR) using specific oligonucleotide primers. PCR products were visualized via agarose gel electrophoresis, cycle-sequenced using d-Rhodamine chemistry, and fractionated on an ABI 377 automated sequencer. Editing of nucleotide sequences and assembly of contig sequences were performed using the computer program Sequencher 3.1 (Gene Codes Corporation, Ann Arbor, MI). Sequences were aligned with POY (Gladstein and Wheeler, 1999) using change/gap cost ratios of 1:2, 1:3, 1:4, 2:5 and 3:5 to test the sensitivity of the phylogenetic conclusions to possible alignment ambiguity. If POY produced multiple, equally parsimonious alignments, each alignment was used in tree reconstruction. In addition, an alignment was performed by eye, and the more variable regions were manually removed from the data set. Trees were rooted to Amphizoa, 30 multiple random addition sequences followed by TBR branch swapping were performed under the parsimony criterion using the computer program NONA (Goloboff, 1994.). Bremer support values were calculated in NONA and bootstrap values were calculated in PAUP* 4.0 (Swofford, 2000). A consensus topology, combining the results of the POY runs under all parameter combinations, was also calculated.
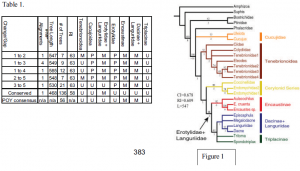
The number of most parsimonious (MP) alignments found, tree length, number of MP trees, retention index (RI), and the groupings of the beetle lineages of interest for each analysis are summarized in table 1. The topology generated with a nucleotide change/gap ratio of 1:2 is presented in figure 1. The topologies produced under the different alignment parameter values are overall very similar. Increasing the change/gap ratio increases the number of MP trees, resulting in a less resolved strict consensus topology. The conserved alignment and the POY consensus analysis produced topologies with the least resolution. These data support the paraphyly of Cucujoidea, with Tenebrionoidea placed within a subset of cucujoid taxa. Tenebrionoidea is generally monophyletic, as is the Cerylonid series. Erotylidae is consistently paraphyletic due to the placement of Languriidae as sister group to Dacne. These data support the monophyly of the subfamily Encaustinae, but consistently produce an unresolved Triplacinae.
Though the taxon sampling in this preliminary analysis is very limited, these results suggest that 18S rDNA appears to be informative at deciphering deeper level relationships within Erotylidae and Cucujoidea. This analysis includes exemplars from only a portion of erotylid subfamilies, and inclusion of additional taxa may help better resolve relationships among these taxa. In particular, this analysis does not include representatives of the Erotylinae and the enigmatic Microsternus. It is clear that additional taxa and genetic markers will provide further insight into the diversification and evolution of this most pleasing of beetle groups.
Working on this research project has been an invaluable experience for me. I presented this research in National Scientific meetings (Entomological Society of America annual meetings) in the form of a research poster. This poster later won first place in the College of Biology and Agriculture’s research poster contest at BYU. The knowledge, skills, and experience I gained during this project will set the stage for the rest of my career.
References Cited
- Boyle, W.W. 1956. A Revision of the Erotylidae north of Mexico (Coleoptera). Buletin of the American Museum of Natural History, 110 (2): 67-142.
- Gene Codes Corporation, Ann Arbor, MI.
- Gladstein, D.S., W.C. Wheeler, 1999. POY. American Museum of Natural History.
- Goloboff, P.A. 1994. NONA [Computer software and documentation]. Tucuman, Argentina: Fundacion e Instituo Miguel Lillo.
- Swofford, D.L. 2000. PAUP*. Phylogenetic Analysis Using Parsimony (*and other methods). version 4.0b4a. Sinauer Associates, Sunderland, MA.