Jennifer Waters and Dr. Mikel R Stevens, Plant and Animal Science
Bromus tectorum (cheatgrass), an invasive winter annual weed, displaces native vegetation, invades crops, and fuels rangeland fires across approximately 40 million hectares of the Intermountain West. Most attempts to control the weed have been unsuccessful, leading to a search for a biological control agent. Ustilago bullata, head smut, is a natural species-specific, fungal pathogen of cheatgrass. Despite apparent phenotypic (physical) uniformity in cheatgrass and head smut, susceptibility and virulence, respectively, are variable. These findings led to the hypothesis that genetic variability for susceptibility/pathogenicity exists in the host and pathogen.
Smut samples were collected from four diverse environmental areas within the Intermountain West. Smut samples from at least 100 infected plants from each area were bulked. I was given 12 sporidial isolates from each area, which I analyzed with AFLPs. AFLPs were chosen as the analysis system for this project because of their ability to detect many polymorphisms. AFLPs are designed to create thousands to hundreds of thousands, depending on the genome investigated, of genomic fragments. These can then be selectively viewed by use of a specific primer combination. Metaphorically, each of these fragments becomes ‘a place we’ve looked in the genome’ and can be numerically expressed as such. AFLPs are also highly repeatable. Once the technique is mastered, experimental results are repeatable both within and between laboratories. This quality substantiates the validity of the results produced through AFLP and, therefore, confidence in their reliability.
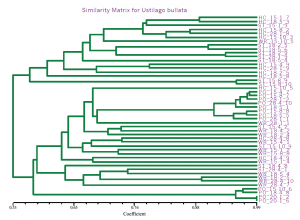
Fifteen primer combinations yielded 1150 total bands of which 150 bands were polymorphic (13.5%). This is a relatively low amount of polymorphism. Analysis, with the SAHN module of the NtSys statistical analysis package as well as the neighbor joining and bootstrap methods of the PAUP 3.0 program, showed consistent segregation of one geographically isolated population into two sub-populations with the remaining three populations showing no distinct patterns (Fig. 1). These results are consistent with the geographic seclusion of this lone isolate set.
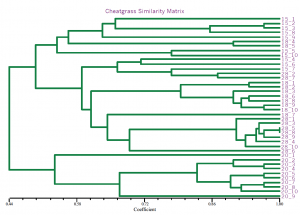
The identification of a genetically isolated strain also interested me in investigating the segregation patterns of cheatgrass AFLPs. AFLPs were performed on 40 cheatgrass lines (10 from each population). Similar segregation patterns were apparent, with the same isolated population segregating independently (Fig. 2). This finding supports the hypothesis that the strain of smut in the geographic region was introduced concurrently with the cheatgrass in that region.
The head smut polymorphism data was also used to generate a quantitative representation of the polymorphism between individuals. Jacob Durrant, a colleague in the lab, developed a program which gave pair-wise comparisons of each individuals to the others and output a percent polymorphism per pair. This type of information would be ideal for choosing parents for a genetic map segregating population for maximum number of possible genetic markers.
Further investigations may reveal additional relationships between observational and molecular data. Given this information, strains of head smut that may be more effective biological controls of specific genotypes of cheatgrass may be developed. The control of cheatgrass should reduce the frequency of rangeland fires and facilitate the successful reintroduction of native vegetation.
Fig. 1 NtSys output for head smut populations. Note clustering of individuals from area ‘PO’.
Fig. 2 NtSys output for cheatgrass populations. Note clustering of individuals from area ’20’.