Steven Woolley and Dr Keith Crandall, Integrative Biology
In my quest to evaluate the efficiency and accuracy of currently used phylogenetic network reconstruction methods, I have learned several valuable lessons applicable to nearly any research endeavor. I have successfully elucidated some of the conditions that affect the accuracy of two popular software programs for intraspecific phylogenetic inference. Below, I will give a brief background, followed by a description of the specific methods that I used in my evaluations. Finally, I will describe and validate the conclusions that I have drawn from my research, and discuss some things that I would have done differently and will do in the future.
Background
Phylogenetic inference is an important biological tool. Phylogenies (or genealogies) allow the study of ancestral relationships within a group of organisms. Traditional methods used to infer and represent phylogenies have been generally limited to bifurcating trees. However, there are many important groups of organisms that are inappropriately represented by such a tree. Some examples of this are viruses (due to rapid evolution, recombination, and other complex ancestral relationships), as well as samples of organisms belonging to the same species or even the same population. This fact has led to several new techniques for representing and inferring intraspecific phylogenies.1 Since the algorithms for reconstructing phylogenetic networks are relatively recent developments, little or nothing had been done to compare the popular methods with each other, or evaluate their strengths and weaknesses. This was the goal of my research project.
Methods
In order to empirically compare different methods for reconstructing network phylogenies, it was necessary to find relevant data to use as input to the software. Since there are very few datasets with known histories, I chose to simulate the data for my evaluations. This required simulation software capable of modeling realistically the processes of molecular evolution. This was done using simulation software created by David Posada,2 which I personally modified for this purpose. Once the data were simulated under a variety of conditions and parameters (16 sets of 1000 histories including DNA sequence data), I needed to convert the data into the appropriate format for use in the software that I analyzed. Initially, I chose six different software packages to evaluate.
Once the data had been evaluated using the selected methods, I needed to convert their output into a common format from which I could compare and contrast the results generally. Additionally, I was faced with the difficult question of how to compare the results. Initially, I considered using a scoring metric known as maximum likelihood. However, as this metric is difficult, if not impossible to use in a phylogenetic network setting in which the network contains a cycle (reticulation), or where the oldest ancestor (the root) is not known, I chose a different set of measurements to compare the results. The measurements that I chose to use were: Robinson/Foulds score,3 Branch score,4 and ambiguity (measured by the number of unique trees contained in each network).
Each of the above measures required that the networks be represented as an enumeration of all possible trees that span the network when combined. I used an algorithm and software from a group of Japanese researchers to perform this task.5 I used the mean and variance of the above three scores to compare the results of two methods on five datasets consisting of 1000 histories each. (See figure 1) The set of trees contained within each network were then compared to the true tree that was simulated.
Conclusions
I simulated 16 sets of data by varying the mutation rate, DNA sequence length, number of sequences, and the evolutionary model by which the data were simulated. As the project progressed, I became aware that those sets of data that were simulated with higher rates of mutation were difficult or impossible to evaluate using some of the methods under study. Additionally, if the networks inferred by an algorithm were highly ambiguous, the number of trees extracted from them was prohibitively large. For this reason, I was only able to evaluate two methods using five sets of data. Nonetheless, some of the results show a definite advantage of one method over the other. (See figure 1)
One important lesson that I learned was that it is vital to perform a meticulous search of research that has been done when beginning a new project. I spent several weeks attempting to implement the algorithm described by Shioura et al,5 before finding their implementation of it. Additionally, in performing the simulations and comparisons, I would have been better off producing a small sample set at the beginning in order to test the entire process, before creating a massive sum of data that couldn’t be used. Aside from these important lessons, I have also learned much from programming the software, interacting with faculty, and reading related literature.
References
- Posada, D., K.A. Crandall. 2001. Intraspecific phylogenetics: Trees grafting into networks. Trends in Ecology and Evolution 16:1, 37-45.
- Posada, D. Coalsim source code and personal correspondence.
- Robinson, D.F., and L.R. Foulds. 1981. Comparison of phylogenetic trees. Mathematical Biosciences 53, 131-147.
- Kuhner, M.K., and J. Felsenstein. 1994. A simulation comparison of phylogeny algorithms under equal and unequal evolutionary rates. Molecular Biology and Evolution 11:459-468.
- Shioura, A., A. Tamura, and T. Uno. 1997. An Optimal Algorithm for Scanning All Spanning Trees of Undirected Graphs. SIAM Journal on Computing 26:3, 678-692.
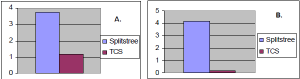
Figure 1. Comparison of SplitsTree with TCS (2 network phylogeny programs). A. TCS demonstrated
less ambiguity. B. SplitsTree varied more in branch score than TCS, on average for one dataset.
