Ermira Pazolli and Dr. Alan Harker, Microbiology and Molecular Biology
Plasmids are genetic entities residing in the cell. They can vary in size and genetic content. There are alternate views about plasmids, but the consensus one is that they are genetic elements that are transferred from cell to cell, thus conferring resistance or performing some other function advantageous to the cell. The primary purpose of this study was to investigate the behavior of plasmids as individuals ‘forced’ to cooperate under selective pressure. In the first phase, we had to extract the plasmids in large quantities and construct two similar models. Due to time constraint and other unforeseen obstacles, we have been able to complete only the first phase.
However, research is progressing and we are currently working on completing the second phase. We used two commercially available plasmids: pEGFP and pEYFP. These plasmids are pUC19 derivates (a high copy plasmid) that contain an antibiotic marker—ampicillin—and fluorescent proteins, green and yellow respectively. These two features made it easier for the plasmids to be detected and followed through time. Initially we had to electroporate the plasmids into an Escherichia coli background. The JM109 strain was selected because it was RecA deficient (it is important for future research when cooperation is studied) and it had good growth properties. Subsequently, the plasmids were extracted using standard extraction protocols. pEGFP had a higher yield at the end of the procedure. This could be explained by some inherent characteristic of the plasmid itself, since both plasmids were treated the same.
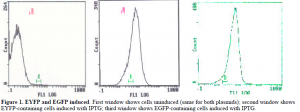
In the next phase, we investigated the fluorescence of the two plasmids. Fluorescence was critical for detection when the two plasmids would be electroporated simultaneously into the same cell in the second phase of the project. We had to detect individual fluorescence first using a Coulter Epics flow cytometer. The genes for the fluorescent proteins were under the influence of the Lac promoter, which can be induced by IPTG to stronger expression. We grew the plasmid-containing cells in MMO medium containing case amino acids. The cells were grown overnight and then IPTG was added, with additional 2-hour incubation. The results of the flow cytometry analysis are shown below in Figure 1.
Both plasmid populations were highly induced by IPTG, and the fluorescence was detectable at high levels. This will allow us to determine the identity of the plasmids in the single-plasmid populations and in double plasmids populations.
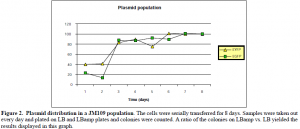
Our next objective was to see the behavior of these plasmids in the cell. We wanted to determine their relationship with the cell population over time. These preliminary data will help us to understand the ecology of these plasmids. Both plasmids were electroporated into cells and they were incubated in LB medium at 37OC, without the presence of any antibiotic. The cultures were serially transferred for a period of eight days. There was no selective pressure for the presence of the plasmid. This experiment determined the natural relationship of the plasmids with their hosts.
The graph shows that the plasmid population reaches a certain level and then it plateaus. We don’t know if this is the case every time or if this was a random event. From other trials (not shown in this report), it appears that the plasmid population plateaus at a level that is different every time. More trials need to be conducted in order to understand the pattern if such a pattern exists. Currently we are setting up a chemostat so that we can follow the plasmid population under more reproducible conditions. We have started the second phase of the project by building the similar plasmid constructs. In order to follow the plasmid population we will have to use two types of specific markers: antibiotic and fluorescent markers. Specific fluorescent markers are present in each plasmid as it was shown above.
We have worked with the antibiotic markers so that each plasmid has a specific marker. The cloning of the antibiotic marker presented challenges that have delayed us in our work. After several months, we succeeded in replacing the ampicillin gene with the kanamycin gene (cloned from a commercial vector) in pEGFP. Even though the plasmids have roughly the same construct they are unique in fluorescence proteins and antibiotic resistance. This will allow us to investigate the behavior of these two plasmids when electroporated simultaneously in the cells. Two outcomes are possible: plasmids can reside in the same cell with one plasmid overtaking the other every time, or the ratio of the plasmids can be different every time due to random events. Our future research will show how these two high-copy plasmids behave.