Theodore L. Oliphant and Dr. Keith A. Crandall, Microbiology
Introduction
This project’s purpose is to determine sites of positive selection in the gp120 envelope protein of the Human Immunodeficiency Virus (HIV). Positive selection is defined as the fixation of changes in amino acid composition in a population. A data set containing the envelope sequence from patients who contracted HIV while participating in a phase 3 efficacy trial of the AIDSVAXTM HIV vaccine was obtained from Vaxgen, the vaccine’s developer. This data is being compared with data sets taken from Genbank to identify sites along the gp120 protein which have mutated, to allow the virus to escape the antibodies generated by the vaccine. Unfortunately, the data set is so large the analyses remain ongoing. However, two separate data sets taken from Genbank were analyzed and yielded interesting results.
Methods
The two data sets consisted of 68 and 79 taxa. The analysis of the data was performed with the following methods and programs. The sequences were first aligned using the program ClustalX (4). Using MacClade 4.05 the alignments were fine tuned and the correct reading frames were determined. The codon numbering was then standardized for each alignment using a lab strain sequence HXB2. In the event that the HXB2 sequence did not have an amino acid at a certain position, the position was numbered by the previous codon’s number in addition to a letter of the alphabet. The data was then entered into PAUP 4.0b10 (3) where a model of evolution was determined using the program ModelTest 3.06 (2). Using the selected model of evolution a neighbor-joining tree was generated for each data set. The data and tree were then entered into TreeSAAP (5), a program that detects positive selection by comparing each sequence to a generated ancestral sequence at each node of the tree. TreeSAAP compares changes in amino acids using 31 physicochemical properties. Each change is assigned a category with 1 being the least amount of change and 8 the most dramatic. TreeSAAP also identifies the individual properties that are undergoing positive selection at a certain category by comparing the ratio of observed substitution events for a particular category to the expected number of events. If a ratio for a specific category is higher than the overall ratio, positive selection is occurring. The program Gblocks (1) was also used to determine ambiguous regions in each alignment.
Results
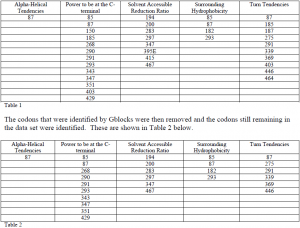
TreeSAAP identified five properties that are undergoing positive selection in relation to category 6, 7, or 8. These are; alpha-helical tendencies, power to be at the C-terminal, solvent accessible reduction ratio, surrounding hydrophobicity, and turn tendencies. These properties are undergoing positive selection at category 8, 7, 7, 7, and 6 and 7 respectively. The individual codons undergoing this selection were then determined and the codons appearing in both data sets are listed in Table 1 below.
Discussion
While the data set consisting of the vaccinated individuals has yet to be fully analyzed by TreeSAAP, interesting points can be drawn from the other data sets. Codons 85 and 87 are implicated as undergoing positive selection in 3 of the 5 properties. These changes as well as the others should be mapped onto the 3-D structure of the gp-120 protein. This would give a more complete picture of what these changes actually mean in terms of the protein. The vaccine data should also be compared to these two control groups.
References
- Castresana, J. Selection of Conserved Blocks from Multiple Alignments for their use in Phylogenetic Analysis. Molecular Biology and Evolution. 2000. 17: 540-552.
- Posoda, David. Keith A. Crandall. Modeltest: Testing the Model of DNA Substitution. Bioinformatics. 1998. 14: 817-818.
- Swafford, D. L. PAUP 4.0b10. Phylogenetic Analysis Using Parsimony and Other Methods. 2002. Sinauer Assoc. Sunderland, MA.
- Thompson, J.D., T.J. Gibson, F. Plewniak, F. Jeanmougin, and D.G. Higgins. The ClustalX Windows Interface: Flexible Strategies for Multiple Sequence Alignment aided by Quality Analysis Tools. Nucleic Acids Research. 1997. 25: 4876-4882.
- Woolley, Steve. Justin Johnson, Mathew J. Smith, Keith A. Crandall, and David A. McClellan. TreeSAAP: Selection on Amino Acid Properties Using Phylogenetic Trees.