Ryan G. Christensen and Dr. David McClellan, Integrative Biology
The nuclei of most extant organisms utilize the universal or standard genetic code. Other codes (e.g. vertebrate, invertebrate, echinoderm, ascidian and insect mitochondrial codes) are quite similar to the universal code (Osawa 1995). However, there are at least 1084 possible genetic codes, each of which encode 20 amino acids and at least one termination signal (Judson 1999). Why then has the standard genetic code come to dominate virtually all life on earth? Theories abound; there are two main schools of thought regarding this question. Some, such as Crick (1968), hold that the universal genetic code came to dominance largely through happenstance. Most theorists, however, assert that the universal code was selected for over the myriad other possible codes because it is the most robust. For instance, Maeshino and Kimura (1998) maintain that the standard genetic code buffers the effects of the most harmful mutations while still permitting mutations to accrue so that evolution and adaptation can occur. This is due to the structure of the genetic code. Codons are arranged so that the most frequent mutations cause no or only slight amino acid changes, while the most harmful and radical amino acid changes are only caused by mutations that occur less frequently.
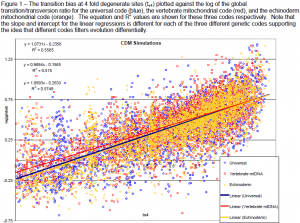
In order to test the hypothesis that different genetic codes filter evolution differently due to their structure, I generated many thousands of simulated DNA sequence data sets (Yang 1996) and calculated the proportion of transitions at 4 fold degenerate sites (ts4) (McClellan 2000) and the global transition/transversion ratio (global(s/v)) for the universal, vertebrate mitochondrial and echinoderm mitochondrial genetic codes. I then plotted these thousand of data points and performed a linear regression.
My results lend support to the idea that the transition bias at 4 fold degenerate sites is highly correlated with the global transition/transversion ratio. However, for the three codes shown here (Fig.1), it is also clear that the same transition bias (ts4) results in a different global s/v ratio, depending on the genetic code. Thus, changing the structure of a genetic code significantly patterns of molecular evolution.
While these results are encouraging and interesting, more work remains to be done. One of the goals of these simulations was to probe the relationship between ts4 and global(s/v) and to express this relationship mathematically if possible. However, the evolver algorithm used to produce the simulated DNA data sets remains a black box in many respects. Thus it is difficult to make conclusions based on out put from this algorithm. The model of molecular evolution that it employs is not entirely clear. Furthermore, evolver is not able to model non-universal genetic codes completely. The frequency of stop codons can be set to zero, but there is no way to change which codons are synonymous and which are nonsynonmous.
Evolver remains a standard tool for researchers studying molecular evolution, but due in part to the difficulties experienced during the course of this study, a new piece of software was developed by David McClellan’s lab. This software, entitled SymSeq, uses a well defined stochastic model of molecular evolution to generate simulated data sets. It allows the user to model different genetic codes precisely. It also provides detailed information regarding all attempted and fixed mutations allowing an investigator to explore the effects of saturation. SymSeq was only recently completed after about a year of development. I hope to continue investigating how genetic code structure shapes molecular evolution using this new software.
References
- Crick F.H.C. 1968. The origin of the genetic code. J. Mol. Biol. 38, 367–379.
- Judson, O. P. and Haydon, D. 1999. The Genetic Code: What Is It Good For? An Analysis of the Effects of Selection Pressures on Genetic Codes. J. Mol. Evol. 49, 539–550.
- Maeshiro, T and Kimura, M. 1998. The role of robustness and changeability on the origin and evolution of genetic codes. Proc. Natl. Acad. Sci. USA 95, 5088–5093.
- McClellan, D. 2000. The codon-degeneracy model of molecular evolution. J. Mol. Evol. 50, 131- 140.
- Osawa, S. 1995. The evolution of the genetic code. Oxford University Press, Oxford, England.
- Yang, Z. 1996. Phylogenetic analysis using parsimony and likelihood methods. J. Mol. Evol. 42:294-307.