Nyall Robert London, Jr. and Dr. Kim O’Neill, Microbiology and Molecular Biology
Cancer remains a major cause of death around the world today. A major problem with cancer is metastasis or the spread of cancerous cells from the tumor to a secondary location. This occurs when cancer cells break away from a primary tumor, invade the tissue, and travel through the bloodstream or lymphatic system to other parts of the body. The aim of this study was to develop a cost-effective in vivo model through which metastasis could be explored in more depth.
Chicken Embryos were chosen to be used as an animal model for several reasons. First, as embryos, they inherently lack a full immune system which allows acceptability of human cancer cells on the chick embryo without a mounted immune reaction. Second, they are an ideal choice because chick embryos are easy to maintain, relatively inexpensive, and have a relatively short incubation period.
In order to further develop and test this model, I chose five cancer cell lines: hepatocellular carcinoma (HepG2), lymphoblast (WTK-1), and three breast adenocarcinoma cell lines (MDA-MB-231, MDA-MB-435, and MCF-7). This selection of cancer cell lines allowed me to ensure that trends seen in tissue specific metastases occurred in a variety of cancers. In order to determine tumor cell invasion, cells were first allowed to incorporate radioactive thymidine into their DNA. This allowed a method through which I could track the cells to various tissues in the chicken embryos. On day eight of chick embryo development, radioactively labeled cancer cells were inoculated on the chorioallontoic membrane (CAM) through a small hole drilled in the shell of the chick embryo. The chicks were then incubated for an additional seven days to permit the cancer cells to metastasize. After these seven days, the chick embryos were sacrificed, various tissues removed, and tissues analyzed for radioactive content using liquid scintillation. The various tissues include the heart, stomach and intestines, lung, brain, and other organs to determine which had the most radioactive labeled cancer cells. This helped us determine which organ or tissue the cancer cells metastasized to most commonly.
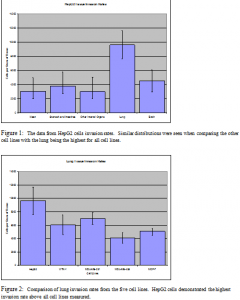
After analyzing the tissue samples, the data was normalized to take into account varying thymidine incorporation rates and tissue sizes. The results I found were very interesting. Figure 1 demonstrates that HepG2 cells predominantly invaded the lung tissue (p value < 0.01). Similar distributions were seen in the other four cell lines. This suggests that the lung was the most common destination to which these five cancer cell lines metastasize. Figure 2 shows the lung tissue rates among the five cell lines with HepG2 demonstrating the highest rate of invasion among all five cell lines (p value < 0.017), suggesting that HepG2 is a valuable cell line to use in models testing cancer cell metastasis.
The project was a success. It allowed me to develop a research idea and accomplish it. In doing so, I also learned about cancer cell invasion. Since completing this project, I have been able to teach this method to other students. The research from this project led to an oral presentation at the 2004 American Society for Microbiology Intermountain Branch Meeting.