Christopher Funk and Dr. Duke Rogers, Integrative Biology
Habromys (crested-tailed mice) is a rodent genus that historically has not been analyzed well enough to be accurately classified. Before 1980, Habromys had been relegated as a subgenus of Peromyscus (deer mice), a closely related genus. As a subgenus, Habromys consisted of only three arboreal species living in the cloud forests of Southern Mexico and Central America (P. simulatus, P. lophurus, and P. lepturus – Hooper and Musser, 1964; Hall, 1981; Musser, 1969). Despite the distinct morphological features of Habromys, it remained as a subgenus due to small sample size and limited morphological analysis. Upon more closely examining its morphology, M. Carleton elevated Habromys to generic status (1980) then later, when he described a new species, H. delicatulus, Carleton proposed Habromys consist of six distinct species (2002). Further, Carleton separated the genus into two morphological groups underscoring the geographic division on the Isthmus of Tehuantepec; H. simulatus, H. delicatulus, and H. chinanteco forming one group on the northwest side of the Isthmus and H. lophurus, H. lepturus, and H. ixtlani forming a group on the southeast side (2002).
While researching, I sought to test the hypothesis that Habromys comprised of six species, including the three recently named species: H. delicatulus, H. chinanteco, and H. ixtlani. I used data that reflected molecular characteristics (based on the cytochrome b gene of mitochondrial DNA) to evaluate the proposed species relationships as described above by Carleton et al. Though my analysis would not seek to concretely confirm Carleton’s proposal, it would provide independent data sets that either strengthened or weakened Carleton’s phylogeny.
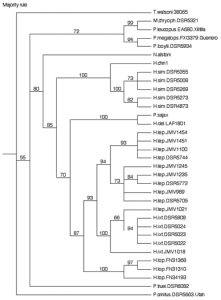
To obtain the aforementioned data I extracted DNA from liver samples, amplified copies of the cytochrome b gene using a PCR machine, purified the PCR product to isolate copies of the target gene, and then submitted this product to BYU’s DNA sequencing center for analysis of the DNA sequence. After constructing sequences for each specimen involved the study, I took the base pair data and performed statistical analyses to produce phylogenetic trees that represented the evolutionary relationship between species based solely on the cytochrome b gene (see figure 1).
Based on the phylogenetic trees of the cytochrome b gene, my data support Carleton’s proposed phylogeny of six independent species. However, it is unclear as to whether H. delicatulus is more closely related to H. simulatus and H. chinanteco grouped in the northwest side or the other three species in the southeast side. My data suggest that H. delicatulus may be more closely related to the southeastern group. To add clarity to the Habromys phylogeny it would be helpful to perform a similar analysis on a nuclear gene from Habromys specimens to bolster Carleton’s proposed phylogeny. Additionally, increasing the sample size of H. chinanteco and H. delicatulus could increase the resolution in the Habromys phylogenetic tree. However, because these species may be extinct, additional specimens may be unavailable.
Literature Cited
- Carleton, M.D. 1980. Phylogenetic relationships in neotomine-peromyscine rodents (Muroidea) and a reappraisal of the dichotomy within New World Cricetinae. Miscellaneous Publications, Museum of Zoology, University of Michigan, 157:1- 146.
- Carleton, M.D. 2002. A new species of Habromys (Muroidea: Sigmodontinae) from Mexico and review of species definitions in the genus, with remarks on diversity patterns among Middle American small mammals restricted to humid montane forests. Proceedings of the Biological Society of Washington, 115(3):488-533 .
- Hall, E.R. 1981. The Mammals of North America. Second ed., John Wiley & Sons, New York.
- Hooper, E. T., and G. G. Musser. 1964. Notes on the classification of the rodent genus Peromyscus. Occasional Papers of the Museum of Zoology, University of Michigan, 635:1-13.
- Musser, G.G. 1969. Notes on Peromyscus (Muridae) of Mexico and Central America. American Museum Novitates, no. 2357, pp. 10-20.