Tom Beckstead and Dr. Jack W. Sites Jr. Department of Biology
Abstract
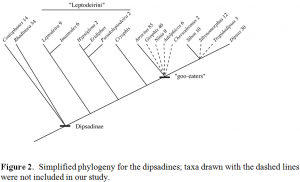
Snakes of the Neotropical subfamily Dipsadinae are poorly known from both ecological and phylogenetic perspectives, despite unique feeding adaptations and bizarre diets. Some of these snakes specialize on snails and slugs (ecologists refer to them as “goo eaters”), and they have a unique dentition and jaw structure that permits them to extract gastropod snails from their shells. Others have enlarged rear fangs, are mildly venomous, and prey on ectothermal vertebrates such as frogs, frog eggs, and lizards (Fig. 1). The most complete phylogenetic study done to date is that of Mulcahy (2007); his study sampled many (but not all) of the species in this clade and constructed a molecular phylogeny based on several mitochondrial DNA (mtDNA) gene regions. Mulcahy (2007) found these two groups of snakes did not form reciprocally monophyletic groups. Instead, the data supported the hypothesis of an ancestral condition of Dipsadines being rear-fanged, mildly venomous, with a derived goo-eater clade (Fig. 2). However, because animal mtDNA normally does not undergo recombination (and therefore independent segregation) typical of genes in the nucleus, all mtDNA genes are effectively linked and inherited as a single unit. Thus while the hypothesis presented by Mulcahy is strongly supported, it is a “single gene tree,” and for many reasons the mtDNA gene tree may not match the true species tree (Degnan and Rosenberg, 2006). This past semester I have been conducting a follow up on Dr. Mulcahy’s previous study by collecting sequences for several nuclear gene regions for the same specimens used in his 2007 study.
Methods
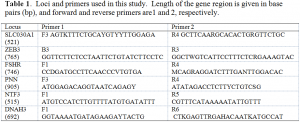
The total genomic DNA was extracted from liver samples of each specimen (all specimens have been catalogued as museum vouchers in professionally curated research collections), and is stored at -80 degrees C in the Sites lab. We used proteinase K cell lysis digestion to extract the DNA (Palumbi, 1996), and then ran polymerase chain reactions (PCR) with different primer combinations (Table 1) to amplify target gene regions. The profiles for each PCR will consist of hot and cold temperatures for replication, and each reaction will contain a forward and reverse primer, in a buffer solution containing dNTPs, Taq polymerase, MgCl (the co-enzyme necessary for PCR) and water. Sequences were obtained for both strands from each specimen. The sequence reactions were then cleaned with Sephadex, edited, translated to amino acids to validate the sequence, aligned, and subjected to several statistical analyses. As I continue working closely with Drs. Sites and Mulcahy, I will be learning how to do the phylogenetic analysis of the sequence data that we obtained through out the semester.
Some preliminary maximum parsimony (MP) analyses were performed on the dataset. The MP analyses were conducted using a heuristic search algorithm with 100 random additions, using tree bisection-reconnection branch swapping. Bootstrap analyses were also performed. The bootstrap analysis is the measure of statistical support. This test was run 100 times with 25 random addition-sequences per replicate.
Results
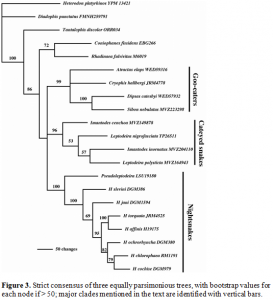
Based on the MP analyses described above, we found that there were three most parsimonious trees with slight differences. A strict consensus of these trees is shown in Figure 3, which demonstrates what all three trees had in common. In the strict consensus tree there is one polytomy of significance. In the first tree the cat-eyed snakes formed a clade with the nightsnakes. In the second two trees the cat-eyed snakes were sister to the goo eaters. The consensus tree shows this as an unresolved polytomy (Fig. 3).
Discussion
Our results based on nuclear sequence data provide weak support for the cat-eyed snakes as sister to the goo-eater clade, contra to the mtDNA results of Mulcahy (2007). However, if these relationships hold up, this does not change Mulcahy’s original hypothesis that ancestral condition of the Dipsadines are rear fanged, and the more derived members are goo-eaters. These new data only re-arrange the relationships among the basal members of the Dipsadinae. As I continue working with Drs. Sites and Mulcahy we will strengthen the hypothesis by collecting more data. We will concatenate the mtDNA data with our newly collected nuclear data and conduct more rigorous analyses when these data are complete, which will hopefully resolve the polytomy in our current topology. I will present results of this work in February 2009 at the Utah Undergraduate Research Symposium at Westminster College, and work with Drs. Sites and Mulcahy to prepare a manuscript for publication. I sincerely appreciate support from the ORCA for my project.