Daniel Tandberg and Dr. Merrill J. Christensen: Nutrition, Dietetics, and Food Science
Introduction
According to the American Cancer Society prostate cancer is the second most commonly diagnosed cancer, as well as the second most common cause of cancer death in the United States. The trace mineral and essential nutrient selenium has been established as a promising chemopreventive element for prostate cancer. Plant foods are the best source of selenium; however, the selenium content of plants varies significantly based on the soil in which it was grown in. This study attempted to provide further insight into the molecular mechanism by which selenium effects cancer development. Specifically we looked at the expression of the Nuclear Factor-kB (NF-kB) and the Androgen Receptor (AR), both of which are transcriptions factors that have a significant impact on the development of prostate cancer. Originally we had planned to use prostate tissue from previously sacrificed Noble rats as our test specimens. However, time and resource constraints instead required us to use cultured prostate cancer cells (LNCaP and 22Rv1) as our specimens.
Methods
Cell Culture: LNCaP and 22Rv1 human prostate cancer cells were maintained in media supplemented with fetal bovine serum and passaged upon reaching confluency. After 72 hours the media was replaced with the treatment media which included various concentrations of selenium in the form of methanoselenic acid (.05 μM control, .3 μM, 5 μM). Treatment lengths were either 48 or 72 hours.
Quantitative RT-PCR: RT-PCR allows researchers to quantify the expression of genes (levels of RNA). Total RNA was isolated from the treated cells with the Qiagen RNeasy kit. The single stranded RNA was converted to double stranded cDNA through reverse transcription. Real-Time PCR was performed using the Roche LightCycler with primer specific annealing temperatures. Primers for 18s, AR, p65, and p50 were used. Prior to using them in the LightCycler they were tested and their optimal annealing temperatures were determined using the RoboCycler and gel electrophoresis. 18s rRNA was used to normalize final concentrations.
Western Blots: Western blots can be used to analyze relative protein levels. Total protein was isolated after the treatment period. Next, the proteins were separated using gel electrophoresis and transferred to a PVDF membrane. The membrane was then incubated with the AR primary antibody followed by a HRP-conjugated secondary antibody. After incubation in a detection reagent the protein was visualized using film. The probing and detection was repeated with the same membrane for the protein Actin, which serves as a control.
Results
RT-PCR: LNCaP and 22Rv1 cells were treated with either .05 μM or 5.0 μM MSA for 72 hours. Four runs with three replicates each were run through the LightCycler. The LNCaP results are shown at right. Selenium significantly reduced the expression of AR and p50 in LNCaP cells. However, selenium also enhanced p65 expression. 22Rv1 cells had a similar reduction in AR and p50 expression, 76.3% and 76.1%, respectively. P65 expression was 86% of the control.
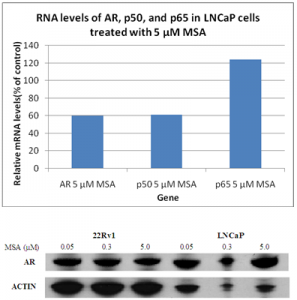
Western Blot: LNCaP and 22 Rv1 cells were treated with either .05 μM, .3 μM, or 5 μM selenium for 48 hours. The results at right are from the second western blot we performed. While the results are not perfect we were pleased to get distinct bands for both AR and Actin. Continued work is being performed in the lab to perfect the western blot protocol. The densitometry analysis of the blot did not indicate any significant reductions in protein levels relative to the control. However, especially with the LNCaP cells, these results are likely highly influenced by human error in preparing the protein samples and loading them into the gel.
Discussion
As with most research a significant amount of time with this project was spent learning and perfecting protocols. While at times this was a frustrating process, through trial and error we were able to produce reproducible results with RT-PCR and improve on our western blot protocol. These experiences required me to develop a stronger understanding of the underlying molecular biology principles upon which they worked. As such I learned at a greater depth than I otherwise would have been able to in a standard classroom
In this particular study selenium in the form of MSA inhibited the expression of the AR and NF- kB genes. This was demonstrated through RT-PCR. While numerous mechanisms likely account for selenium’s anti-cancer properties, it is possible that the effect of selenium on these genes is a significant mechanism. Continued studies are warranted on selenium and prostate cancer to further elucidate these molecular mechanisms.
