Keri G. Dockter, Derek S. O’Neil, David L. Price, John Scott, and Mikel R. Stevens, Plant and Wildlife Sciences
Tomato spotted wilt virus (TSWV), vectored by several thrips species, is the causal agent of devastating tomato crop losses in many areas of the world. Recently, field tests have demonstrated that there are TSWV isolates that overcome the resistance gene Sw-5, derived from Lycopersicon peruvianum L2. However, Sw-7, a new source of TSWV resistance, has been introgressed from L. chilense Dun1. This new gene has demonstrated field resistance to various isolates of this disease including greenhouse trials utilizing isolates that overcome Sw-51. In order to determine the genetic location of Sw-7, we screened over 200 SSR and InDel molecular markers from across the tomato genome3. We used the homozygous (Sw-7/Sw-7) CK12 line and one of the susceptible (Sw-7+/Sw-7+) recurrent backcross parents (Fla 7482B) along with six BC1 and F2 plants segregating for Sw-7. The results of this screening suggested that Sw-7 resided near SSR20 on chromosome 12. To confirm the linkage of Sw-7 and SSR20 we tested this SSR marker and others on chromosome 12 using 100 BC1 and F2 progeny segregating for Sw-7. Additionally, we screened 47 lines segregating for Sw-7 under high natural TSWV field pressure conditions in northern Florida4. As a result of these studies, we have narrowed the location of the L. chilense-derived DNA with Sw-7. This region resides between markers C2_At4g16710 (40.0 cM) and CT189 (59.0 cM), according to the “Tomato EXPEN-2000” map found on the Sol Genomics Network. Our efforts are now focused on determining the precise position of Sw-7 in relationship with the markers available in this region and the AFLP markers previously reported.
Materials and Methods
• F2 seed was grown in the greenhouse and leaf tissue was collected, regardless of disease response. We expected a normal Mendelian ratio among these plants.
• We extracted genomic DNA from leaf tissue from the parent lines (CK12 and Fla 7482B) and from 94 F2 progeny
• Polymorphic markers identified with a parental study were tested on the F2 population on a 1% GenePure agarose gel and then scored in an excel sheet.
• Fragments of interest that weren’t polymorphic but gave a single band during electrophoresis were tested with Sanger sequencing for SNPs (single nucleotide polymorphisms) and InDels (insertion/deletions), and then scored.
• Short-length polymorphisms were tested on a polyacrylamide gel using the Li-Cor 4300 DNA Analyzer.
Results
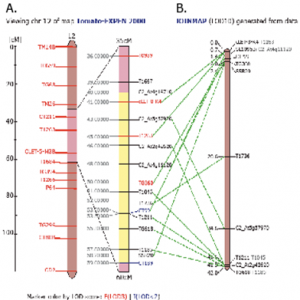
The purpose of this study was to map this region with Sw-7 present, and no deliberate selection pressure. We tested over 200 SSR (simple sequence repeat) markers on DNA from the parent lines and found a section of wild L. chilense DNA containing Sw-7 on chromosome 123. Our data suggests that Sw-7 is between 40 cM (C2_At4g16710) and 59 cM (CT189), according to the “Tomato EXPEN-2000” map. There are 46 markers found in this region, with over half of them above 55 cM. However, some were not easy to work with; some seemed to correlate to different areas of the genome. These particular markers were not included in the results of this study. Furthermore, while this introgression segregates 100% with Sw-7 resistance, chi-square analysis of the data in JoinMap 45 suggests a different ordering of the markers for this population. C2_At5g57970, C2_At2g42620, T1045, T1736, T1211, TG618, and T1185 were all significantly different from the expected Mendelian ratio. JoinMap 4 generated this map as the only possible ordering for these markers5.
Discussion
JoinMap 45 generated the most probable order for these markers in our population. The candy stripe pattern witnessed in Table 1A makes the current order very improbable—especially when an inversion may also be involved. This in part explains why some of the marker loci do not represent any heterozygotes. Additionally, our chi-square analysis has placed these particular markers at a very significant distance from the others.
The next step in this project will involve a comparison of data with the F2 seed that was sent to Florida1. This will also include the back-crosses that were made to develop a desirable but resistant cultivar. This may further elucidate the location of the Sw-7 gene within the area of wild L. chilense DNA.
References
- Canady MA, Stevens MR, Barineau MS (2001) Euphytica 117:19-25
- Gordillo LF, Stevens MR, Millard MA, Geary B (2008) Plant Dis 92:694-7904
- Price DL, Memmott FD, Scott JW (2007) TGC Report 57:35-36
- Scott JW, Stevens MR, Olson SM (2005) TGC Report 55:40-41
- Van Ooijen, J.W. JoinMap 4 (2006)® Kyazma B.U., Wageningen, Netherlands
