Camille Finlinson Porter and Dr. Byron Adams, Biology
Meloidogyne nematodes are agricultural pests that cause approximately 5% of global crop loss each year. In order to develop more effective control strategies, a clearer image of how the many species of Meloidogyne are related is needed. A phylogenetic tree is a hypothesis of the relationship between different species. Phylogenetic trees allow us to see how closely related species are by comparing a few genes from many species. Although numerous phylogenies of the genus have been constructed in the past, none of them summarize relationships across the entire genus because of incompatibility between taxa and genetic datasets. Each species that has been studied has had different genes sequenced. This makes it difficult to compare species relationships. The objective of my research was to merge several phylogenies for different genes in the Meloidogyne genus and make a comprehensive phylogenetic tree using a supertree approach. A supertree is the summary the relationships of input trees called subtrees.
Preparation for this project included taking a graduate class – Phylogenetic Systematics (Biology 640). The knowledge I gained from taking the class was essential in order for me to complete the project.
I completed this project in two phases. The first was to make a supertree out of previously published trees; the second was to make a supertree of trees that I created. We used trees published by DeLey, Tigano, Landa, Tenente, Castillo, and Lunt to create a supertree (Fig. 1). The relationships in the trees they published were combined to make our tree. This tree is resolved, but it could be inaccurate. The problem with using other people’s trees is that each tree was created with different parameters. What is considered optimal in one tree would not be in another. This is why it was necessary to create my own trees- each of the gene trees that I created was made using the same conditions.
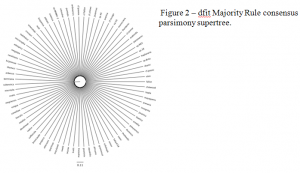
The data for the second phase came from Genbank, an online database of DNA. All sequences for every gene available for Meloidogyne was taken from the database. This analysis included more species than the first. I made a separate tree for each gene. In order to make a supertree, the relationships of the gene trees were combined to make one big tree. (Fig. 2). Unfortunately, the tree that I created is completely unresolved; the relationships between species are wholly unknown. The gene trees disagreed with each other so much that summarizing them gave us no information. More work should be done to see if there is a way to make the tree more resolved.
This work was published in 2009 in the book Root-Knot Nematodes (CABI Publishing) in chapter 5 – Molecular Taxonomy and Phylogeny. I also presented it at the annual meeting of the Society of Nematologists in Vermont this July.