Chad C. Lowe and Dr. William R. McCleary, Microbiology
Within the cellular environment, the processing of essential nutrients required for cellular growth and metabolism are tightly regulated. When these essential nutrients become limited in the cells surroundings, cellular stress responses are induced. This enables the cell to maintain life in less than ideal environmental conditions. Recent studies with the bacteria Escherichia coli have shown a mechanism for phosphate uptake into the cell under such conditions. When phosphate is scarce, a two-component regulatory system is activated to facilitate phosphate transport (5). This system, called the Pho System, consists of a sensor kinease (PhoR) and an activator (PhoB). When phosphate is limited in the environment, PhoR becomes autophosphorylated and in turn phosphorylates PhoB (3). A conformational change is induced as a result of phosphorylation. PhoB forms a dimer and then recognizes and binds to a consensus sequence called the pho box; transcription of stress response genes is then enhanced (1). These genes code for proteins and enzymes that help the cell “scavenge” for phosphate. Alkaline phosphatase (AP) is an enzyme produced by one of these genes; cellular AP levels increase as phosphate is reduced in the environment (2). AP cleaves bound phosphate in the periplasmic space of the cell so that it can be transported into the cytoplasm.
In an effort to understand the structure and function relationships of PhoB, previous research in the McCleary lab has generated random mutations in the phoB gene. Of particular interest are mutations in the N-terminal regulatory domain; mutations in the C-terminal DNA binding domain of the protein generally cause a negative phenotype (i.e. AP is not expressed) whereas mutations in the N-terminal regulatory domain of the protein cause a constitutively active (always active) phenotype. It has been previously shown that amino acids at certain positions (9,10,11, and 54) conform to create the active site of the protein (6). A recently isolated mutation in PhoB at amino acid position 11, which changes a glutamic acid (E) to a basic lysine (K) residue, was found to show constitutive activity; i.e. AP is expressed at both high and low levels of extracellular phosphate. This current research project investigated some of the possible mechanisms that cause this protein to be constitutively active.
The following methods were used to first purify the mutant protein and then explore it’s affinity for the pho box:
Protein Purification. The specific mutation, E11K, was introduced into an over-expression plasmid vector, pT7-phoB, by site-directed mutagenesis (SDM). Primers containing the specific mutation were used in a polymerase chain reaction (PCR) to introduce the mutation onto the DNA template, after which the original methylated DNA was digested with the enzyme Dpn-1 (DNA formed in the PCR reaction is non-methylated). The genetic sequence of the induced mutant phoB was then verified using a Licor-4000 automated sequencer.
The pT7-phoB(E11K) plasmid contains an inducible operon upstream from the phoB gene. This plasmid was transformed into an appropriate strain, after which a one liter culture was grown at 37EC. When the optical density was found to be 0.1 at 600nm, isopropyl-b-D-thiogalctopyranoside (IPTG), at a final concentration of 0.2mM, was added to induce high levels of E11K protein. Protein overexpression was then verified by SDS-PAGE. At this point it was found that the protein was insoluble in the standard buffer used previously in wild-type PhoB over-expression. The insoluble protein was hydrolyzed in a solution of 8M urea, after which renaturization was accomplished by dialysis in a buffer
glycerol, and 10mM 2-mercaptoethanol, for a final concentration of about 10 μL/mL. The protein was then concentrated using a centrifugal membrane filter to an approximate concentration of 0.3 mg/ml.
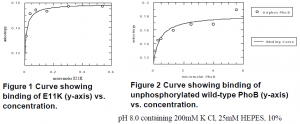
DNA Binding Assays. The affinity of E11K for the pho box was tested using fluorescence anisotropy. This technique measures the rotation of a flourescent-labeled molecules in solution. E11K protein at increasing concentrations was introduced into a solution containing a short segment of labeled DNA with pho box. The relative rotation of this molecule was compared to that of unbound DNA; increased binding leads to a higher anisotropy (or slower rotation).
These preliminary results show that unphosphorylated E11K binds DNA with an approximate 30-fold affinity over unphosphorylated wild-type PhoB. These results therefore demonstrate the constitutive property of this protein. Future research will focus on verifying the dimer state of the mutant protein by size exclusion chromatography and preparing these results for publication. Further purification and concentration of the protein will lead to crystallization studies, which will elucidate the mechanism of the constitutive properties of E11K.
References
- Makino, K., Shinagawa, H., Amemura, M., Nakata, A. 1986. Nucleotide sequence of the phoB gene, the positive regulatory gene for the phosphate regulon of Escherichia coli K-12. J. Mol. Biol. 190, 37-44.
- Makino, K., Shinagawa, H., Amemura, M., Kimura, S., Nakata, A. 1988. Regulation of the phosphate regulon of Escherichia coli: activation of pstS transcription by PhoB in vitro. J. Mol. Biol. 203, 85-95.
- Makino, K., Shinagawa, H., Amemura, M., Kawamoto, T., Yamada, M., Nakata, A. 1989. Signal transduction in the phosphate regulon of Escherichia coli involves phosphotransfer between PhoR and PhoB proteins. J. Mol. Biol. 210, 551-559.
- Volz, K. 1993. Structural conservation in the CheY superfamily. Biochem. 32(44), 11741- 11753.
- Wanner, B. L. 1993. Gene regulation by phosphate in enteric bacteria. J. Cell. Biochem. 51, 47-54.
- Zundel, C.J., Capener, D.C., McCleary, W.R. 1998. Analysis of the conserved acidic residues in the regulatory domain of PhoB. FEBS Letters. 441, 242-246.
