Crystal Komm and Dr. Keith Crandall, Zoology
Phylogenetic analyses are used to examine evolutionary relationships among crayfish species at different taxonomic levels. Nuclear genes provide genetic information from independent loci which, when analyzed, can be compared to mitochondrial DNA sequences. Astacin is a nuclear gene encoding a digestive protein in crayfish. This was the focus of this study’s analyses. In this study we had the following five aims:
1) to determine whether astacin is a single of multiple copy gene
2) to compare our astacin sequences to the Astacus astacus sequence obtained from Genbank through examining the genetic structure of the astacin gene from exon 3 through exon 5
3) to perform a phylogenetic analysis of astacin sequences collected during this study
4) compare the phylogeny obtained for astacin sequences with those produced for 16S mitochondrial DNA and morphology.
Methods
Crayfish astacin sequences were collected from 3 Orconectes species and 2 Procambarus species representing the Cambaridae crayfish family. Samples from Austropodomobius and Astacus represented the Astacidae family. Sequences were compared at various taxonomic levels from within individuals to among families. Sequences were obtained through DNA extraction, amplification via Polymerase Chain Reaction, cloning, and automated sequencing according to standard laboratory procedure.
Analysis
Sequences were blast searched on Genbank to verify that they were crayfish astacin. Alignment was accomplished via Clustal X (Thompson et al. 1997). Modeltest 3.0 (Posada & Crandall 1998) was used to calculate likelihood scores for 56 different models of evolutions. A maximum likelihood analysis was executed in PAUP* (Swofford 1999) with 100 heuristic searches and implimenting the appropriate model of evolution. Confidence in nodes was reassessed using 100 bootstrap replicates (Felsenstein 1985). DNA sequences were also translated to protein using MacClade (Maddison and Maddison 1999) for intron-exon analysis.
Results
397 bases of astacin sequence were obtained from seven crayfish individuals. Sequences cloned from the same individual differed by a maximum of 2 base pairs, with the exceptions of O. virilis, which differed by up to 4 base pairs. Our sequences differed from the Astacus astacus from Genbank, which contained a fourth intron. Our sequences had a large fourth exon that was equivalent to exon 4 and 5 in A. astacus. Translation into amino acid sequence revealed no stop codons in the exons (protein coding regions). The combined exons contained 61 unambiguous base pair changes, 38 of which were synonymous (no amino acid change) substitutions and the other 23 were non-synonymous substituions (amino acid change). The ratio of non-synonymous to synonymous substitutions was 0.1634. Modeltest determined the data set to be most compatible with the HKY + gamma model of evolution (Hasegawa et al. 1985). The phylogenetic reconstruction generated by PAUP* is shown below in Figure 1 with bootstrap values indicated on the lines. All nodes were well supported with high bootstrap values.
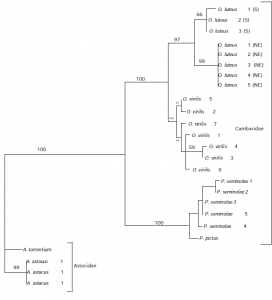
Conclusion
The lack of variation among clones suggests that astacin is a single copy gene. This precludes astacin evolution within an individual from being examined. However, higher levels of variation in the O. virilis clones may be attributed to the beginning of a duplication event. Levels of variation increased with the taxonomic hierarchy sampled and indicate that the astacin gene will be useful in phylogenetic analyses at a number of taxonomic levels including the population level and higher. The phylogenetic tree obtained for astacin was the same as that obtained for the mitochondrial 16S gene region (Crandall & Fitzpatrick 1996) and for morphological data (Hobbs 1988) suggesting it is a reliable indicator of relationships.
References
- Crandall KA, Fitzpatrick JF (1996) Crayfish molecular systematics: using a combination of procedures to estimate phylogeny. Systematic Biology 45:1-26.
- Felsenstein J (1985) Confidence limits on phylogenies: an approach using the Bootstrap. Evolution 39:783-791.
- Hasegawa M, Kishino H, Yano T (1985) Dating of the molecular clock splitting by mitochondrialDNA. Molecular Biology and Evolution 22:160-174.
- Hobbs HH (1988) Crayfish distribution, adaptive radiation and evolution. In Freshwater crayfish: biology, management and exploitation (ed. DM Holdich & RS Lowery), pp. 52-82. Timber Press, Oregon.
- Maddison WP, Maddison DR (1999) MacClade 4.0: Analysis of phylogeny and characterevolution. Sinauer Associates, Sunderland, MA
- Swofford DL (1999) PAUP*4.0: Phylogenetic Analysis using Parsimony. Sinauer Associates, Sunderland, MA.
- Thompson JP, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The ClustalX windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Research 24:2875-82.
