Chinn-Woan Shih and Richard Robison, Department of Microbiology and Molecular Biology
Main Text
As stated in the proposal, my focus has been with three bacterial species of Burkholderia: mallei, pseudomallei, and thailandensis. Burkholderia mallei is the causative agent of glanders, an abscess-forming infection that is predominantly found in the equine population and is capable of being transmitted to humans from infected animals. There is documentation that B. mallei was used as a biowarfare agent in World War I (Gee et al., 2003).
B. pseudomallei, a soil pathogen, is the causative agent of meliodosis, a tuberculosis-like disease that is endemic in regions of southeast Asia and northern Australia. B. thailandensis is nonpathogenic for humans and animals but displays phenotypic characteristics that make it appear similar to B. pseudomallei by routine diagnosis tests (Thilbault et al., 2004).
Because these three species of Burkholderia are very similar, molecular methods of rapid detection and differentiation have been a topic of recent research. If a test was available that clearly differentiated all three species, one could quickly detect the presence of these bacteria form soil, animal, or human samples. With such a test, detection of these pathogens, especially in the early stages of infection, would prevent further development of the disease, which would significantly decrease the morbidity and mortality rates in humans. It appears that as increasing numbers of diagnostic laboratories in endemic regions of Asia are established, and of increased travel, these diseases will be recognized increasingly during the next decade, thus furthering the significance of developing such a test (White, 2003).
Quantitative real-time PCR (qPCR) has revolutionized diagnostic testing due to its ability in detecting specific genetic signatures of organisms of interest (Saunders). To date, no multiplex qPCR assay has been developed that can detect and differentiate between these three species in a single-tube format; this is the purpose of my research. My research group was able to develop primers that seem to be specific for B. mallei & pseudomallei. We did not have a successful primer set that was specific for B. thailandensis; the bulk of my research has been to develop one. After testing through several primers and analyzing data, we were able to develop a primer set that may work.
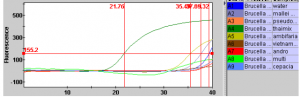
Initial testing of this B. thailandensis primer set was performed using SYBR-green (double-stranded DNA binding dye) before purchasing the TaqMan probes (detection of a specific PCR product). In figure 1, specificity for the B. thailandensis cocktail mixture was found to be specific compared to other Burkholderia species. A few near-neighbor species were detected, but at later cycles and with significantly smaller fluorescence. Given this preliminary data, I decided changing the PCR cycles to 35 instead of 40 would make the primers more specific. After further tests of the primer using SYBR-green, my research group was confident that purchasing the TaqMan probe would be our next stage of specificity testing. The probe that we developed also showed some specificity for B. multivorans. We were somewhat concerned with this, but this was the closest we have been to creating a primer set that is specific for our species.
Figure 1-B. thailandensis (green) has spiked at an earlier cycle than all other species of Burkholderia
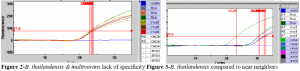
Also, from previous publications in qPCR assays, we have found that the TaqMan probes tend to yield the same results as SYBR-green as long as the primers are specific for the species of interest; this was the case with the B. thailandensis primers. Temperature sensitivity tests for the primers and TaqMan probe proved to be unsuccessful. The annealing/extension temperature ideal for SYBR-green denatured the probe. This proved to be a problem because the forward primer could only be specific if it was at its ideal temperature; a temperature that the probe could not withstand. As a result, B. multivorans fluorescence was about the same as B. thailandensis (Figure 2). Also, the species of Burkholderia was also showing amplification at cycles around 30 (Figure 3). As mentioned earlier, specificity for B. thailandensis proved to be more specific if the cycle parameters were around 35. Making the parameters to 30 would not provide enough data for series dilution testing (testing for the primer set sensitivity).

I was not able to complete my goals according to the proposed timeline. As of right now, it seems like the B. thailandensis primer/probe set could possibly be specific. I have proposed three approaches to determine its specificity:
1. Perform a multiplex test: by doing this, detection sensitivity of all the primers would decrease which may decrease specificity for B. multivorans.
2. Quantify the DNA isolates and dilute to the same concentration: the isolates that have been tested could significantly vary in the amount of DNA. If this is the case, the near neighbors may no longer be a problem.
3. Make modifications to the forward primer: the primary goal is to lower the annealing/extension temperature without compromising specificity for the species.
References
- Gee, J. E., Sacchi, C. T., Glass, M. B., De, B. K., Weyant, R. S., Levett, P. N., Whitney, A. M., Hoffmaster, A. R. & Popovic, T. (2003). Use of 16S rRNA gene sequencing for rapid identification and differentiation of Burkholderia pseudomallei and B. mallei. J Clin Microbiol. 41, 4647-4654.
- Saunders, N. A. “An Introduction to Real-Time PCR.” Edwards, Kirstin, Logan, Julie, Saunders, Nick. Real-Time PCR: An Essential Guide. London: horizon bioscience, 2004. 1-12.
- Thibault, F. M., Valade, E. & Vidal, D. R. (2004). Identification and discrimination of Burkholderia pseudomallei, B. mallei, and B. thailandensis by real-time PCR targeting type III secretion system genes. J Clin Microbiol 42, 5871-5874.
- White, N. J. (2003). Melioidosis. The Lancet. 361,1715-1722.