Mark Rowan and Dr. Bradford K. Berges, Microbiology & Molecular Biology
Kaposi’s Sarcoma Herpesvirus (KSHV) is a prevalent disease around the world. Some Sub- Saharan African countries have infection levels as high as 40 percent. Usually the effects of this disease are not fatal unless the infected person has already been infected with HIV or has some other immunodeficiency. If both of these problems exist, deadly cancers can form within the infected person1. Studies have shown that people with HIV are 200,000 times more likely to have Kaposi’s sarcoma (a cancer caused by KSHV) than those not infected. Until now, no competent animal model existed to study KSHV pathogenesis.
In our laboratory, we use Rag2-/-γc-/- which are born devoid of an adaptive immune system, and do not produce murine T or B cell responses. Three days after birth, we engraft the mice with human hematopoietic stem cells, which serve as a source of both the cells infected by KSHV and a human immune response to the virus. These humanized mice have previously been shown to be infected with the highly related Epstein-Barr Virus, and showed early signs of cancer. Our hypothesis was that Rag2-/-γc-/- humanized mice2 could be used to study KSHV infection and disease. Since the formation of a latent3 (no active viral replication) or lytic (active viral replication) viral infection plays a critical role in the disease state of the patient, my proposal focused on determining which type of infection (lytic or latent) existed in our mice.
KSHV has eighty genes, five of which are exclusively latent genes. I planned to quantify the expression level of six genes (Orf65, RTA, Lana, VIL-6, V-cyclin, Gapdh). Some of these genes are exclusively lytic, and others are exclusively latent. This would allow our lab to know which type of KSHV infection is present in the humanized mice. There wouldn’t be enough infected cells in the blood, so I planned on collecting cells from the lymph nodes, spleen, bone marrow, and peritoneal cavity. All of these areas contain KSHV target cells. We planned on using uninfected mice as negative controls, and KSHV positive tumors which expressed our genes of interest as a positive control. After collecting these cells, I was going to extract the RNA. Then, using gene-specific primers, I planned on constructing cDNA from their respective RNA templates. Then, using TaqMan probes, I was going to use quantitative PCR to quantify the expression of the six genes I chose to study.
As it often happens in science, however, my project did not go forward as planned due to genetic impurities in our mouse colony. Somehow our colony was contaminated with immunocompetent mice and this prevented us from starting my project, as well as any other project in our lab. We have spent the last six month purging our colony, and just recently began to create humanized mice once more.
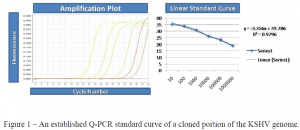
I did as much work on my project as I could without having the actual humanized mice. I finished the cloning of the six genes (with the help of another student in the lab) and made all the preparations to create standard curves for the gene expression of these six genes. Then, I turned my focus to an experiment that involved reactivation of KSHV in BCBL-1 cells.
Reactivating KSHV in BCBL-1 cells allowed us to get everything ready so that when we do have a sufficient number of humanized mice, all we have to do is collect the RNA and analyze it via Q-PCR. Instead of collecting KSHV RNA from humanized mice, we were collecting KSHV RNA from BCBL-1 cells, which made for a similar experiment. This also helped us optimize everything for the forthcoming KSHV infection of humanized mice. We will not have a lot of humanized mouse RNA with which we can work, so it is imperative that we have all of the experimental conditions set beforehand.
To begin the experiment I defrosted and then cultured BCBL-1 cells until I had 3 T-75 flasks full of cells. I saved one flask for normal culturing, and used the other two for the experiment. I combined the two experimental flasks, spun down the media, and counted the cells to ensure there were an equal amount of cells in each flask. I counted 20.9 million cells, and used 11 million for the experiment, 5.5 million in each experimental flask. 5.5 million cells were reactivated and the other 5.5 million cells were used as a non-reactivated control.
I added 1mm sodium butyrate and 50 ng/ml of 12-O-Tetradecanoylphorbol-13-acetate to stimulate KSHV reactivation in the reactivated flask. Then, I let the cells culture overnight until they were ready for RNA extraction. I then used an RNA extraction kit to extract the RNA from the reactivated and non-reactivated cells. After the RNA isolation I then reverse transcribed the extracted RNA. Once we have all the standard curves ready these RNA levels will be able to be compared to the standard curve levels.
Now that our mouse colony has been purified, we should have this project completed in a few months. I recently graduated from BYU, and I will be attending the UCLA School of Dentistry in the fall. Therefore, I will not be around for the completion of our project, but I feel I have done everything possible to ensure it is finished quickly and smoothly.