Michael Ahlborn and Dr. Brian Poole, MMBIO Dept.
Introduction
Systemic lupus erythematosus, commonly known simply as lupus, is an incurable autoimmune disorder that can cause inflammation and tissue damage in virtually any part of the body. Many studies have shown a correlation between lupus diagnosis and positive infection with Epstein-Barr Virus (EBV). Scientists currently estimate that greater than 90% of the adult US population is infected with this human herpesvirus, probably because it is easily transferred through saliva. Although we do not entirely understand how it contributes to lupus it is apparent there is a link between the two. Other research has shown that EBV may also play a role in infectious mononucleosis, and certain types of cancer, such as Hodgkin’s lymphoma and Burkitt’s lymphoma. It is important to note that many of the people who test positive for EBV infection exhibit no signs or symptoms associated with these illnesses. Genetics and environmental factors, as well as EBV, are believed to play an important role in whether or not a particular disease develops.
Toll-like receptor 7 (TLR7) is an important innate immunity receptor protein whose activation is also linked to lupus. When this receptor is stimulated in normal B cells it causes an increased expression of interferon regulatory factor 7 (IRF7), which is a crucial regulator of type I interferons in pathogenic infections. Previous studies have demonstrated that EBV infection is linked to decreased expression and activation of IRF7 through sumoylation (posttranslational modification). While performing experiments related to the IRF5 protein we noticed that several other proteins including IRF7 were downregulated in EBV positive cells after TLR7 stimulation by imiquimod. This observation attracted our interest in IRF7 and led to the hypothesis upon which this study is based.
Methodology
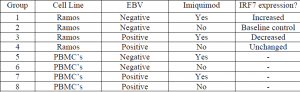
Ramos (Human Burkitt’s lymphoma cell line) cells and peripheral blood mononuclear cells (PBMC’s) were the two types of cells I originally intended to use among several treatment categories. However, due to a variety of difficulties I had with using PBMC’s, I have not been able to get clear results relating to these cells yet. I also had some challenges at first getting our EBV positive cells to express the green fluorescent protein. Nonetheless, I was able to complete much of the project following the guidelines that were originally proposed. Cell lines were grown up using standard protocols and RPMI media. After that, extraction of RNA, reverse transcription of RNA to cDNA, and real-time PCR were performed according to recommendations supplied by Dr. Poole and Daniel Clark (graduate student).
Results
Discussion
The goal of this project was to observe IRF7 expression in EBV infected and non-infected B cells treated with imiquimod (stimulates TLR7). My hypothesis was that IRF7 expression would decrease in EBV positive cells that have been treated with imiquimod. At this time it appears that my results fail to disprove the hypothesis. However, these results are only preliminary and there still remains a significant amount of work to be done. While completing this project I ran into several challenges. One big one was that I have not been able to work successfully with the PBMC cell lines. I expect that continued patience and practice will help me master my technique and be able to do so in the future. Another unanticipated challenge was the time required to grow significant quantities of Ramos cells. I am used to rapidly dividing cells and wasn’t prepared for the weeks required for Ramos cells to grow. An additional setback was that after completing some of the procedures, I learned of a better/more efficient way of doing them with a lower chance of experimental error, so I look forward to redoing some of them in order to get more precise results.
Conclusion
This project has been a great learning experience and helped me understand how to manage the complexity of different procedures with very different timetables. I learned how to separate B cells from a fresh sample of blood and how to perform RNA extractions as well as reverse transcribe RNA to cDNA and was also able to use my previous knowledge about polymerase chain reactions (PCR) in my experiments. Although I encountered many setbacks, I still believe future work would make it possible to fully test my hypothesis. I am extremely grateful to have been mentored by Dr. Poole and to work in his lab. I am also thankful for the funding I received from the ORCA office in order to continue my education in this excellent learning environment.