David Peterson and Dr. John S. K. Kauwe, Biology
Introduction
Alzheimer’s disease (AD) is the most common neurodegenerative disease, affecting more than 4.5 million people in the US. Genetic studies of AD have previously identified mutations in three genes (APP, PSEN1 and PSEN2) and polymorphism in APOE as risk factors. These findings have led to a better understanding of the underlying disease mechanisms. However, half of all AD cases have no known genetic risk factors for disease. Most studies are designed to identify variants associated with risk or age at onset, but rarely cover other important facets of AD, such as disease progression or duration. Previously, Cruchaga et al. identified several single-nucleotide polymorphisms (SNPs), located in the genes encoding the regulatory subunit of the protein phosphatase 2B (PPP3R1, rs1868402), and the microtubule-associated protein tau (MAPT, rs3785883) genes, that are associated with CSF tau levels. They further found that rs1868402 and rs3785883 were associated with an increased rate of progression of AD. In this study we attempted to support these associations by genotyping these two SNPs in an independent sample of 92 Alzheimer’s disease cases from the Dementia Progression Study and testing them for association with the rate of progression of Alzheimer’s disease as measured by the Clinical Dementia Rating sum of boxes (CDR-SB).
Materials & Methods
We genotyped rs1868402 and rs3785883 in 92 subjects with incident AD from the Dementia Progression Study using custom Taqman® Assays. Subjects had a mean initial age of 84.5 years, 45% were male, and 47% were APOE ε4 carriers. Subjects were selected to have a Clinical Dementia Rating (CDR) of less than 1 at their initial clinical assessment and CDR sum of boxes (CDR-SB) data for at least two time points. Subjects had an average of 4 assessments and average time from first to final assessment was 3.18 years. To determine the significant covariates age at first assessment, apolipoprotein epsilon 4 (APOE ε4) presence/absence, and gender were tested for association with change in CDR-SB over time (rate of progression) using PROC MIXED in SAS version 9.2. Only gender showed significant association and was therefore included as a covariate in the final models to test rs1868402, rs3785883, and the combined genotypes of rs1868402 and rs3785883 for association with rate of progression.
Results
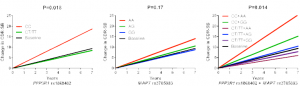
For rs1868402 a recessive genetic model was the best fit with minor allele homozygotes, “CC”, showing significantly increased rate of progression (p=0.018; “CC”=2.67 boxes per year (bpy) and “CT/TT”=1.21; figure 1). Rs3785883 did not show significant association, but showed a trend toward faster progression with each copy of the minor allele, “A” (p=0.17; “GG”=1.25 bpy, “AG”=1.5, “AA”=2.0; figure 1). The model incorporating genotypes from both markers in combination yielded significant association with individuals that carry risk genotypes at both markers progressing at 3.6 bpy (p=0.014; figure 1).
Conclusion
Our analyses support the association found by previous studies that rs1868402 and the combined genotypes of rs1868402 and rs3785883 are associated with the rate of progression of AD. Although our results with rs3785883 alone were not statistically significant, the direction of the effect observed in our data is consistent with the report. The data reported here provides a better understanding of the genetic variability that influences the rate of progression of Alzheimer’s disease and could provide novel insights into possible preventative and therapeutic strategies.
Academic Outcome
We presented the results of this study at the International Conference on Alzheimer’s Disease in Paris, France in July 2011. I was the presenting author on the poster and had the opportunity to defend our findings to clinicians and other researchers at the conference. Funds from my ORCA grant, supplemented by a MEG, and various other grants from the NIH enabled me to travel to Paris for this conference. I am currently drafting a final manuscript of this study, which we hope will be published by the summer.
Works Cited
- Hayden KM, Zandi PP, Lyketsos CG, Tschanz JT, Norton MC, Khachaturian AS, Pieper CF, Welsh-Bohmer KA, Breitner JCS (2005) Apolipoprotein E Genotype and Mortality: Findings from the Cache County Study. Journal of the American Geriatrics Society 53:935–942
- Hebert LE, Scherr PA, Bienias JL, Bennett DA, Evans DA (2003) Alzheimer’s disease in the US population: prevalence estimates using the 2000 census. Arch Neurol 60:1119-1122
- Harold D, Abraham R, Hollingworth P, Sims R, Gerrish A, Hamshere ML, Pahwa JS, et al. (2009) Genome-wide association study identifies variants at CLU and PICALM associated with Alzheimer’s disease. Nat Genet 41:1088- 1093
- Cruchaga C, Kauwe JSK, Mayo K, Spiegel N, Bertelsen S, Nowotny P, Shah AR, Abraham R, Hollingworth P, Harold D, Owen MM, Williams J, Lovestone S, Peskind ER, Li G, Leverenz JB, Galasko D, Morris JC, Fagan AM, Holtzman DM, Goate AM (2010) SNPs Associated with Cerebrospinal Fluid Phospho-Tau Levels Influence Rate of Decline in Alzheimer’s Disease. PLoS genetics 6:e1001101
- Kauwe JS, Cruchaga C, Mayo K, Fenoglio C, Bertelsen S, Nowotny P, Galimberti D, Scarpini E, Morris JC, Fagan AM, Holtzman DM, Goate AM (2008) Variation in MAPT is associated with cerebrospinal fluid tau levels in the presence of amyloid-beta deposition. Proc Natl Acad Sci U S A 105:8050-8054
- Norton MC, Piercy KW, Rabins PV, Green RC, Breitner JC, Ostbye T, Corcoran C, Welsh-Bohmer KA, Lyketsos CG, Tschanz JT (2009) Caregiver-recipient closeness and symptom progression in Alzheimer’s disease. The Cache County Dementia Progression Study. J Gerontol B Psychol Sci Soc Sci 64(5):560-8.
