Jordan Mackay and Dr. Julianne Grose, Molecular and Microbiology
Many diseases that affect humankind are poorly understood. The medical and scientific communities alike rely heavily on basic scientific research to fuel dissemination of knowledge to said communities for further applied research, including clinical and pharmaceutical efforts. Diabetes is still poorly understood despite its prevalence in today’s society. In lieu of the research that has been done so far, PAS kinase is becoming an increasingly promising subject for research as a possible therapeutic target for diabetes. Studies have shown that PAS kinase is involved in metabolic regulation and mouse experiments have been done which show these results.1 As we learn more about PAS kinase and identify its downstream targets, the more therapeutic targets we will have for treating the adverse effects of type II diabetes. What my research team and I have uncovered in the past few months may prove to have such an impact, pending further research.1
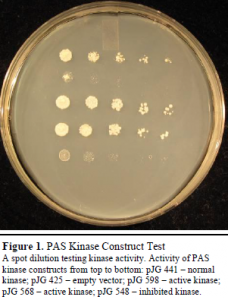
We were able to successfully construct a version of PAS kinase in which the kinase domain is constitutively active (see Figure 1). The construct pJG 598 was chosen for the screening procedure. Compared to screens with pJG 441, the screening with pJG 598 was found to be more successful and generated many more hits. These findings coincide with other studies of PAS kinase structure and function.2
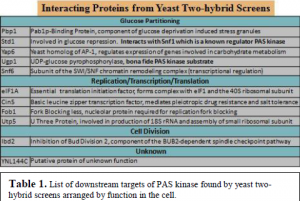
Through repeated screening procedures, we discovered the interactions of PAS kinase with the products of the genes Pbp1, Ibd2, Cin5, Yap6, Snf6, and several other previously uncharacterized yeast genes (see Table 1). These results are very exciting, as many of these genes are involved in the processes of central dogma, i.e. transcription and translation. Each one has to be fully sequenced to ensure the gene is in its entirety and that it is properly fused to the Gal4 activation domain. They also have to be further tested to be proven as bona fide substrates of PAS kinase.
The plasmids harboring our interactor of interest need to be isolated from yeast and independently transformed back into yeast along with proper controls. Many of these genes have undergone such tests two to three times and have proven in this test to be true interacting partners of PAS kinase.
There still remains much to be done on this project to prepare for publication. Namely, there must be in vivo and in vitro proof of not only an interaction between PAS kinase and its substrate, but also proof of phosphorylation or other such events characterizing it as a true substrate-kinase relationship. We plan on performing co-immunoprecipitation assays as well as in vitro and in vivo kinase assays in order to provide the scientific community with information regarding the nature of these interactions. Once complete, we will have obtained enough information to publish our findings.
When my team first began performing these screens and other tests, we encountered a lot of problems with contamination. We also struggled with using Aureobasidin A, a fungal antibiotic, as a reporter. We overcame these problems by developing extremely sterile technique in performing screens as well as choosing to screen with other reporters in place of Aureobasidin, namely adenine and histidine auxotrophies. We presented our preliminary results at the regional ASM conference at Utah State University in April, 2011 in the form of an ASM-style poster. We have since gathered a greater quantity of data to present for the upcoming year, which we will also publish.
Although far from being over, this research opportunity has taught me much about patience, hard work and dedication, and organizational skills that will aid me in my future endeavors in scientific research.
References
- Rutter J, Hao HX. 2007. The role of PAS kinase in regulating energy metabolism. IUBMB Life. 1-29.
- Grose JH, Smith TL, Sabic H, Rutter J. 2007. Yeast PAS kinase coordinates glucose partitioning in response to metabolic and cell integrity signaling. The EMBO Journal; 26(23): 4824-4830.
