Serena Loeb and Dr. Julianne Grose, Microbiology and Molecular Biology
Prevalence of metabolic disease, primarily diabetes, is increasing at a steady rate. This trend has created a need for new treatment options. Researchers discovered that the protein PAS (Per-Arnt-Sim) kinase is involved in mammalian metabolic regulation; showing PAS kinase deficient mice exhibited decreased weight gain, decreased liver triglyceride accumulation, decreased insulin resistance, and an increased metabolic rate. This encouraged the design of human PAS kinase inhibitors. The exact mechanisms of PAS kinase remain unknown. Therefore, the aim of our research was to find substrates of PAS kinase to better assess health risks associated with PAS kinase inhibitors, and provide new potential therapeutic targets for type II diabetes.
We studied the functions of PAS kinase in humans using Saccharomyces cerevisiae as a model because PAS kinase is evolutionarily “conserved from yeast to man” (Hao & Rutter, 2007). The rapid growth rate and genetic pliability of yeast were some of the benefits we encountered while studying PAS kinase in this system. PAS kinase already had one known substrate, UDP-glucose phosphorylase 1 (UGP1). However, kinases typically perform several functions in the cell and the PAS kinase deficient mice showed a wide range of effects, which suggested that PAS kinase had other substrates (Hao & Rutter, 2007). Our project focus was to discover these substrates.
METHODS
To achieve this goal, we used the yeast two-hybrid screen developed by Fields and Song. Yeast two-hybrid screens generally produce the best results when multiple constructs of the protein of interest are used (Geitz, 2006). Therefore, we screened with several constructs to find whether a truncated or full-length construct of PAS kinase would yield more substrates.
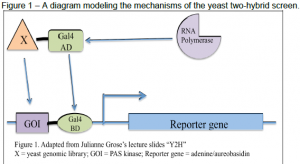
In the yeast two-hybrid screen, Gal4 wild type protein recruits RNA polymerase to allow for transcription of target genes. By breaking Gal4 into two components, the DNA binding domain (BD) and the DNA activating domain (AD), transcription cannot occur unless these domains are brought back together to recruit RNA polymerase and activate the transcription process (Geitz, 2006). See Figure 1. We then bound the AD to the yeast genomic library and the BD to our gene of interest (a construct of PAS kinase). If any of the genes from the yeast genomic library interacted with PAS kinase, we saw growth. We then patched these colonies from the screen and tested them on various media sources for functionality. For those that grew as expected, we isolated the plasmid from the yeast genomic library and sequenced it to identify the plasmid.
We used the full length PAS kinase construct to perform the screen. The PAS kinase construct interacted with the yeast genomic library, thus causing the BD and the AD to interact and recruit RNA polymerase. This triggered transcription and growth. We isolated the genomic library plasmids out of the yeast and sequenced the plasmids. We had a hit—Erv29. We did the same with a truncated construct of PAS kinase, and after sequencing the colonies that grew on our media from that screen we found Ibd2, Pbp1, and YIL108W. Both of these screens also yielded false positives, which showed that our experiment was working.
RESULTS
Erv29 is an ER-associated protein involved in formation of COPII-coated vesicles. These vesicles are required for transport of a subset of secretory proteins into the Golgi, including insulin. Ibd2 is involved in the inhibition of bud division. It interacts with Bfa1 and Bub2p, constituents of the GTPase-activating complex. This complex is involved in mitosis. Pbp1 is a constituent of glucose deprivation induced stress granules. It interacts with Pab, a polyadenylation protein involved in mRNA regulation. YIL108W is an uncharacterized metalloprotease, which is a metal dependent enzyme that begins protein digestion by hydrolysis of peptide bonds (yeastgenome.org). It is likely that these proteins were involved in the results seen in the PAS kinase deficient mice in Jared Rutter’s lab. Further research must be done to discover what other proteins interact with PAS kinase.
Throughout the screening process, we ran into problems with contamination. When this occurred, we checked our reagents for contamination and used greater precaution in experiments. We saw results with the yeast two-hybrid screen, as we expected. We have found multiple interactors from our screens that are very promising. This leads us to believe we are on the right track to finding many more. We presented our research in the form of a poster at the American Society for Microbiology Intermountain Branch Annual Meeting at Weber State University in April of 2011.
REFERENCES
- Hao HX, et al. PAS kinase is required for normal cellular energy balance. PNAS 2007; 104(39)15471.
- Geitz RD. 2006. Yeast Two-hybrid System Screening. Methods in Molecular Biology; 313: 345-371.
- http://yeastgenome.org/. Accessed June 16, 2011.
Acknowledgements:
Dr. Julianne Grose, Steve Sowa, Katie Harris, Colby Haines, Jordan Mackay