Mo Lee and Dr. John S.K. Kauwe, Biology Department
Alzheimer’s disease is the most common form of dementia in the United States. It is a complex neurodegenerative disorder featured by gradual onset and progression of memory loss combined with deficits in executive functioning, language, visuospatial abilities, personality, behavior and self-care. There are basically two types of Alzheimer’s Disease, familial Alzheimer’s disease and late-onset Alzheimer’s Disease respectively.
The pathogenesis of Alzheimer’s Disease still remains largely unknown and the proposed mechanisms are still under much investigation. Recent studies indicate a fast increase in the prevalence of AD unless more effective treatments can be developed. The cost of medical care used by those patients is astronomical because it takes a large amount of effort to assist the patients who will eventually lose all survival skills and die slowly. In this study, we determined to provide more information on the genetic factors that influence YKL40. The goal of this study was to identify the genetic variation that influences cerebrospinal fluid levels of YKL40 and determine how that variation relates to risk and rate of progression of Alzheimer’s disease.
In our studies, we focused on late-onset Alzheimer’s disease. We used a linear regression model in our analyses, and our focus was on the gene called YKL40, which is also called CHI3LI, chitinase 3-like 1. It is shown that there is increased YKL40 gene expression in schizophrenia, autism and Alzheimer’s Disease (Bonneh-Barkay 2010) We have obtained extensive phenotype and genotype data from our collaborators at the Washington University School of Medicine. Quantitative measurements of YKL40 in cerebrospinal fluid were obtained using ELISA in approximately 300 individuals. Individuals were also genotyped using the Illumina OmniExpress chip, providing nearly 1 million genetic markers across the human genome.
This large dataset requires specialized analytic techniques and computational resources. We first uploaded our data to Kauwe lab shared folder on Mary Lou. Plink(Purcell 2007), an online genetic analysis package that has been installed in the super computer, was used to test for association between genotypes at each marker and cerebrospinal fluid levels of YKL40. Analysis was performed using linear regression models. As YKL40 was not normally distributed in these data a parametric statistical approach is inappropriate. To account for this problem we used a non-parametric permutation approach to generate an empirical distribution of p-values. To minimize the amount of false positive associations we used 10^-6 as our p-value cut-off for significance. Markers that show association with YKL40 levels were tested for association with risk and rate of progression of Alzheimer’s disease.
Half way through the project, I realized that we wouldn’t be able to finish according to the timetable as I planned due to the size of our dataset and the complexity of the analyses. My co-worker, David McKean who was a Bioinformatics major, was able to construct a python script that allowed me to perform the analyses in parallel, allowing the project to finish on time.
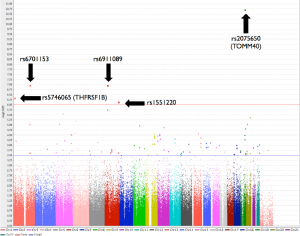
We found significant association with CSF YKL40 levels for rs2075650 (P value=1.96E- 11) in TOMM40, rs6911089 (P value= 1.08E-07), rs6701153 (P value= 1.10E-07), rs5746065 (P value= 4.54E-07) in THFRSF1B, rs1551220 (P value= 7.13E-07).
 I presented our results at a BYU student poster session. I am still currently working on putting together a manuscript for our project. In the future, we need to investigate more on TOMM40, which is a gene that provides significant increased precision in the estimation of age at onset of AD for the patients with APOE3 allele. We expect studies focused on this gene to lead to fruitful results.
I presented our results at a BYU student poster session. I am still currently working on putting together a manuscript for our project. In the future, we need to investigate more on TOMM40, which is a gene that provides significant increased precision in the estimation of age at onset of AD for the patients with APOE3 allele. We expect studies focused on this gene to lead to fruitful results.
