JanaLynn Franke and Dr. Brad Geary, College of Life Sciences, Department of Plant and Wildlife Sciences
Background
Cheatgrass (Bromus tectorum) is one of the leading causes of premature wildfires in the Great Basin and is almost always the establishing species following a fire. Fire has historically been an important part of the Great Basin ecosystem however; disruptions of natural fire regimes have altered diversity of vegetation. Previously the natural fire regime consisted of fires occurring approximately 80 to 100 years apart. The current fire regime has been reduced to as little as 5 percent of that. Cheatgrass germinates and matures quickly, making it extremely competitive with native grasses and will continue to replace native species unless control measures are instituted.
Main Body
Localized areas within the Great Basin have been observed to resist Cheatgrass growth while surrounding areas continue to host Cheatgrass. Within the past few years massive research has been done to try and identify the driving force behind mass Cheatgrass die-off. Due to the patterns of the die-offs it is believed that a pathogen or combination of pathogens is to blame. Owen Baughman, a graduate student from the University of Idaho conducted an experiment that isolated pathogens found in die-off soils. In his experiment, he attempted to prove that these pathogens where behind the monoculture stand failure. Unfortunately, the hypothesis that soil borne pathogens were the causal agents, and the data obtained, did not coincide with one another. Within the last year, my research has shown that the original hypothesis is correct. My research accounted for a couple of variables that were not considered when Owen Baughman did the initial study. In order for a disease to be successful there are four variables that need to be taken into account: environment, host, time and of course the pathogen. In Owens research only two (Host and pathogen) out of the four were established. With the knowledge of how pathogens work, we were able to identify the problem and retest the pathogens and produce successful data that agreed with the original hypothesis.
Material and Methods
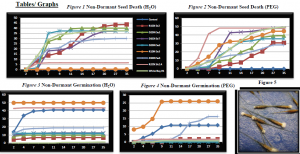
In the Previous experiment done by Owen Baughman Ring molds were used to obtain soil samples located inside the Dead spots in Skull Valley. These were then placed in a greenhouse and Cheatgrass (Bromus Tectorum) seeds planted in each ring mold and were watered regularly. Unfortunately there was no significant decrease in germination.1 The new procedure took into account all aspects (including time and environment) of disease survival: environment, host, time, and pathogen. In the pathogens native environment the water potential was around -1.5. We achieved this by using Polyethelyn glycol (PEG) a chemical that restricts moisture available to the seed. During the observation of the dead spots it was noticed that the seedlings initial growth in the late fall was non-existent meaning, the pathogen must have inoculated and penetrated the seed during the summer months. With these two aspects accounted for we were able to develop an experiment that would ensure the pathogens success. The experiment proceeded as follows. Each fungi was grown out on 1/3 PDA and allowed uninhibited growth for 2 weeks. They were then injured (mycelium removed to stimulate spore production) and let grow for 2 more weeks. At that time the spores were harvested to be used in a liquid inoculum between [1.25*105]-[2.5*105] spores per milliliter, and divided up into two equal parts one for Dormant and one for non-Dormant (the seeds used were previously surface sterilized in 10% bleach and 75% alcohol). The seeds were soaked in the inoculum for 1 minute to allow for inoculation of seeds. Seed were then placed in a petri-dish lined with a bladder that was soaked in either autoclaved H2O or PEG, each dish receiving 50 seeds. There were a total of 4 runs for each pathogen: Dormant H2O, Dormant PEG (-1.5 ѱ), Non-Dormant H2O, Non-Dormant PEG (-1.5 ѱ). These runs were incubated at 25 degrees Celsius (previously determined to be an optimal temperature) for 35 days. At day 7 seeds that were in PEG were then switched to a H2O bladder to simulate the natural environment of summer drought and fall rain storms. Numbers of seeds germinated and seed deaths were collected on days 2, 4, 7, 11, 14, 17, 20, 27, and 35.
Results
Data obtained confirmed the previous hypothesis that soil borne pathogens were the causal agents of Cheatgrass stand failure. There was a marked decrease in seed germination and a marked increase in seed death when seeds were exposed to a negative water potential. This proves that the negative water potential of -1.5 found in the Great basin ecosystem gives an advantage to the pathogen, allowing it to penetrate the seed while the seed is unable to germinate. As a result of the test I believe that it is a combination of these pathogens that are responsible for these exhibited dead spots. Each pathogen has a different killing time within the 35 days. Specific pathogens that are the leading killers are R2CN Sh2 A and D1CN Se7. I used Light microscopy along with a fellow team member who used genetic sequencing to identify these pathogens2. Both techniques worked equally as well and all of the following pathogens were identified as Fusarium. The following data will be published in a paper submitted to Weed Research in January 2012. It will be titled “Exploring the causes and effects of Cheatgrass die-offs in the Great Basin” and Authors listed as such: Owen Baughman, Susan Meyers, Julie Beckstead, and JanaLynn Franke.