Michael Baldwin and Dr. Laura Bridgewater, Department of Microbiology and Molecular Biology
Introduction
Nuclear Gdf5 (nGdf5) and nuclear Bmp2 (nBmp2) are newly discovered proteins (1); however, their function has not been determined. They were first found while studying enhancer elements of the Col11a2 gene, and a yeast-one hybrid screen suggested that they could bind to the D/E enhancer of this gene. Further experiments confirmed the presence of these proteins in the nuclei of cells and determined the specific mechanism by which they were translocated to the nucleus (1). This work led to the hypotheses that they function as transcription factors, binding to regulatory regions of target genes to modulate transcription. Two years ago, I identified a potential consensus DNA binding sequence for the protein nGdf5 using a technique called cyclic amplification and selection of targets (CASTing). The initial intent of this project was to do the same for nBmp2, but I decided that further analysis of nGdf5 was needed to prove that I truly identified a consensus DNA binding sequence for nGdf5.
Objective
The purpose of this project was verify that nGdf5 binds the sequence (T/G)GGTG(G/T)G and to determine its specificity to the four possible variations of this sequence. The results of this project would determine if CASTing should be performed with nBmp2 and the work examining the potential role of nGdf5 and nBmp2 as transcription factors should be continued.
Methods and Results
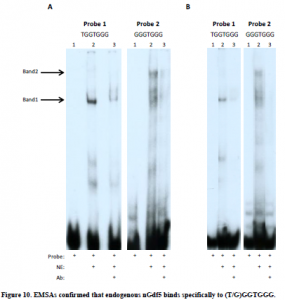
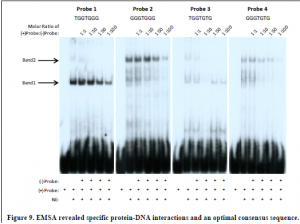
To confirm that nGdf5 binds to the sequence (T/G)GGTG(G/T)G, DNA probes representing each of the four possible binding sequences (TGGTGGG, GGGTGGG, TGGTGTG, and GGGTGTG) were synthesized and utilized in three EMSAs. In the first EMSA, radiolabeled probes were incubated with nuclear extract from non-transfected 10T1/2 cells to see if an endogenous protein would bind to any of these sequences. Increasing amounts of unlabeled probe was included in each binding reaction at a concentration of 1-,10-, 50-, and 100-fold molar excess over the concentration of the radiolabeled probe to act as a specific competitor. Two major bands appeared on the EMSA that were competed away by unlabeled probe, indicating that both of these bands represent specific protein-DNA interactions. Although the same protein-DNA interactions were detected using each DNA probe, the intensity of these interactions were weak with probes 3 and 4. Probe 3 and 4 both have a thymine at position six of the consensus sequence, whereas probe 1 and 2 have a guanine at this position, suggesting that (T/G)GGTGGG is a more optimal consensus sequence (Figure 9). The results seen in the first EMSA demonstrate that proteins in the nuclear extract of 10T1/2 cells bind in a sequence-specific manner to probes 1and 2. To determine if either protein was nGdf5, another EMSA was performed by adding anti-Gdf5 antibody to the binding reactions with probes 1 and 2. Anti-Gdf5 antibody blocked both the protein-probe 1 interactions and the protein-probe 2 interactions, revealing that although the protein-DNA complexes migrate with different electrophoretic mobilities, both contain nGdf5 (Figure 10A). A third EMSA produced the same result, confirming that nGdf5 binds to the consensus DNA binding sequence (T/G)GGTGGG (Figure 10B).
Conclusion
My research demonstrates that nGdf5 binds specifically to the consensus sequence (T/G)GTGGG. Further experiments should be conducted to determine whether nGdf5 and nBmp2 function as transcription factors. Due to their involvement in cartilage and bone formation, understanding their role as transcription factors may provide new information on bone maintenance and cartilage breakdown, and may establish new means for treating arthritis.
Works Cited
- Felin, Jenny E., et al. “Nuclear Variants of Bone Morphogenetic Proteins.” BMC Cell Biology 11:20 (2010).