Alisa Paulsen and Dr. Daniel Fairbanks, Botany and Range Sciences
In the Western United States, cheatgrass (Bromus tectorum, introduced from Eurasia) is often considered to be an undesirable, invasive weed. When cheatgrass is introduced into an area, it can rapidly establish itself as the dominant species, inhibiting reestablishment of native plants, competing with important grain crops such as winter wheat, and increasing the frequency of fire.1,2,3 Over the last one-hundred years, cheatgrass has spread so rapidly that it is now a dominant grass in large areas of the United States.
Because cheatgrass inhabits quite a broad range of ecosystems, the rapid spread of cheatgrass is probably due to multiple introductions. Cheatgrass is an obligate inbreeder, so each unique introduction results in a unique maternal line identical to the parent.4 Using genetic markers, it is possible to fingerprint these lines and determine the extent of each maternal line within and among different populations. We have isolated six such markers and have screened plants from four different populations as a test of the sensitivity of the markers. These markers take advantage of the fact that DNA can be quite different among closely related individuals.
Simple sequence repeats (microsatellites) are a type of highly variable DNA. DNA is composed of combinations of four molecules (nucleotides) commonly referred to as adenine (A), thymine (T), cytosine (C) and guanine (G). Microsatellites are simply places in the DNA where nucleotides, for example C and G, are repeated over and over again, somewhat like a stutter or hiccup. The DNA sequence from one individual may contain six repeats of CG and another individual that seems identical in other ways may contain only five repeats of CG. The advantage to microsatellites is that it is not necessary to determine the exact sequence of each microsatellite in each individual plant; instead, all we have to determine is the exact length of each microsatellite in each individual. If a microsatellite is 12 nucleotides (base pairs, or bp) long in one plant and 10 nucleotides in another, we know they are from different maternal lines.
In order to develop these microsatellite markers, we had to construct a library of cheatgrass DNA. This library contains small pieces of cheatgrass DNA that we inserted into phages using the ZAP Express Predigested Vector Kit (Stratagene). The kit yielded a genomic library of approximately 100,000 different phages (clones). We screened approximately 11,200 clones for microsatellites by attaching the DNA from the clones to nylon membranes and hybridizing the membranes with CA and CT oligonucleotide polymers labeled with DIG (Roche). Positive clones were detected by color detection (NBT/BCIP) and sequenced with a Perkin-Elmer ABI-Prism automated sequencer.
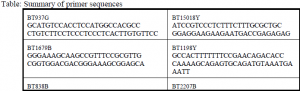
Ten clones each contained six to 18 dinucleotide repeats. In order to detect these microsatellites in each plant, we designed primers (custom pieces of DNA flanking the dinucleotide repeats) to yield amplification products of 87bp to 240bp. Amplification of DNA is performed by mixing Taq polymerase (Promega) with the DNA and allowing it to repeatedly replicate the area flanked by the primers in a process called the polymerase chain reaction, or PCR. We labeled the primers with fluorescent markers for use on the automated sequencer, and optimized PCR conditions on an Eppendorf Mastercycle Gradient thermocycler by varying temperature and magnesium concentration. Final PCR reactions were performed on a Perkin-Elmer 9600 thermocycler.
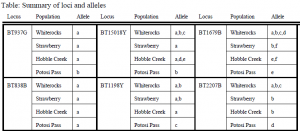
Of 10 loci that contained microsatellites, two had serious sub-banding problems, and two didn=t amplify. We screened 53 individuals from four different cheatgrass populations of the Great Basin: Whiterocks (UT), Strawberry (UT), Potosi Pass (NV), and Hobble Creek (UT). BT838B appeared monomorphic in all populations, and all other loci had from two to six alleles. Individuals from Potosi Pass, a recently invaded high desert, had only one allele of each locus, and those alleles were found almost exclusively in that population. This suggests that the Potosi Pass population was introduced separately from the other lines we identified. Samples from Whiterocks, a desert in western Utah, amplified three alleles, suggesting the existence of three maternal lines. TheStrawberry population seemed to be composed of only two lines. We plan to use these loci to screen a greater sample size from a wider variety of locations.5
References
- Melgoza, G., Nowak, R. S., Tausch, R. J. Oecologia 83:7-13.
- Young, J. A., Evans, R. A. J. Range Mngt. 31:283-289.
- Blackshaw, R. E. Weed Sci. 41:551-556.
- Susan Meyer, USDA Forest Service Shrub Sciences Laboratory; personal communication.
- Acknowledgments: Dr. Craig Coleman, Department of Botany and Range Sciences, Dr. Susan Meyer, USDA Forest Service Shrub Sciences Laboratory.