Chad C. Lowe and Dr. William R. McCleary, Microbiology
Within the prokaryotic cell, the intake of essential nutrients is continuously regulated by intracellular mechanisms. When fundamental nutrients become scarce, cellular stress responses are activated to enable the cell to survive in a harsh environment. Recent studies have shown a two-component regulatory mechanism which enhances Escherichia coli intracellular phosphate uptake in a phosphatepoor environment (5). This system, called the Pho system, consists of a sensor (PhoR) and an activator (PhoB). When environmental phosphate is limited, PhoR becomes autophosphorylated and in turn phosphorylates PhoB (4). The phosphorylated PhoB recognizes and binds to a consensus sequence called the “pho box” and transcription of phosphate uptake and processing genes is enhanced (2). PhoA, or alkaline phosphatase (AP), is an enzyme produced by one of these genes; intracellular levels of PhoA increase as phosphate concentrations decrease in the cellular environment (3). PhoA cleaves bound phosphate in the periplasmic (outer) space of the cell so that it can be transported into the cytoplasm, or inner area of the cell.
Previous research has isolated random mutants in the C-terminal DNA binding domain of PhoB; these mutations inactivated the PhoB protein. This mutagenesis research was conducted in a high copy (i.e. 200 copies per cell) plasmid (a small circular piece of DNA containing the phoB gene). It has been hypothesized that a similar mutagenesis conducted on a low copy (i.e. 1-20 copies per cell) plasmid would more closely resemble the cell in vivo and result in a more even distribution of mutants between the N-terminal and C-terminal PhoB domains. The objective of this research project was to generate phoB mutations and analyze those found specifically in the N-terminal regulatory domain of the protein.
A low copy number plasmid containing the phoB gene was inserted into a cell lacking DNA repair mechanisms. The cells were grown for several generations, allowing random mutations to develop within the plasmid. These plasmids were then harvested and inserted into cells lacking the PhoR and PhoB components of the Pho system. After the cells were grown on minimal media (media containing a limited predetermined amount of phosphate), dyes were sprayed on the plates to test for PhoA activity. Those colonies showing no PhoA activity or abnormal activity were isolated; the phoB gene was then sequenced to determine where exactly the mutation had occurred.
Western blot analysis showed the majority of mutantions produced truncated PhoB protein. One sequenced mutation showed multiple mutations in the phoB region, which could explain why complete PhoB protein was not found. Multiple mutations may have changed the protein configuration such that PhoB was inactivated. One mutant, however, exhibited high levels of PhoA activity at both high and low levels of phosphate, suggesting a constitutively active (always turned on) mutant. This mutant was sequenced (Figure 2); amino acid eleven was found to change a glutamic acid to a basic lysine residue. In other two-component regulatory systems, introduction of a basic amino acid in the acidic active area of the protein produced constitutive activity (5).
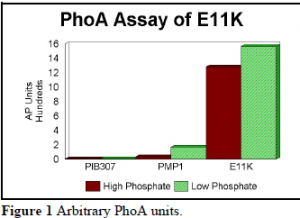
An AP assay was conducted to quantify the relative amount of PhoA produced in the apparently constitutive mutant (Figure 1). The pIB307 plasmid contains no phoB, the PMP1 plasmid contains wildtype phoB, and E11K is the apparent constitutively active mutant.
A recent study of an N-terminal PhoB mutant has shown that a constitutively active mutant is not phosphorylated (6). Protein from the mutant E11K will therefore be purified and phosphorylation assays will be conducted to determine how activation of the Pho system occurs in this mutant. Other assays may be conducted to determine the mutant protein’s affinity for the pho box.
References
- Bourret, R., Hess, J., Simon, M. 1990. Conserved aspartate residues and phosphorylation in signal transduction by the chemotaxis protein CheY. Proc. Natil. Acad. Sci. 87, 41-45.
- Makino, K., Shinagawa, H., Amemura, M., Nakata, A. 1986. Nucleotide sequence of the phoB gene, the positive regulatory gene for the phosphate regulon of Escherichia coli K-12. J. Mol. Biol. 190, 37-44.
- Makino, K., Shinagawa, H., Amemura, M., Kimura, S., Nakata, A. 1988. Regulation of the phosphate regulon of Escherichia coli: activation of pstS transcription by PhoB in vitro. J. Mol. Biol. 203, 85-95.
- Makino, K., Shinagawa, H., Amemura, M., Kawamoto, T., Yamada, M., Nakata, A. 1989. Signal transduction in the phosphate regulon of Escherichia coli involves phosphotransfer between PhoR and PhoB proteins. J. Mol. Biol. 210, 551-559.
- Wanner, B.L. 1993. Gene regulation by phosphate in enteric bacteria. J. Cell. Biochem. 51, 47-54.
- Zundel, C.J., Capener, D.C., McCleary, W.R. 1998. Analysis of the conserved acidic residues in the regulatory domain of PhoB. FEBS Letters. 441, 242-246.