Crystal Komm and Dr. Keith A. Crandall, Zoology
Astacin is a zinc-endopeptidase, originally described as a unique enzyme and first sequenced from the crayfish Astacus astacus (Geier 97). This project aimed to gather several Astacin gene sequences from multiple taxa of crayfish and to submit them to phylogenetic analysis using PAUP (Swofford 98). PAUP would then generate trees displaying the most probable evolutionary relationships among taxa based on variation in the Astacin gene sequences.
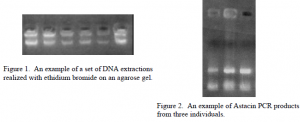
The first step toward obtaining gene sequences required extracting DNA from alcohol preserved and fresh-frozen crayfish tissues. The extraction method used was a standard Proteinase K protocol adapted from Crandall (99). The extracted DNA then served as the template for the Polymerase Chain Reaction (PCR). PCR amplified the Astacin gene region for sequencing.
In order to carry out PCR, previously designed primers, specific to the Astacin region were used. It was created from the Astacus astacus Astacin sequence. A series of PCRs using this set of primers yielded no PCR products. A variety of temperatures and cycle variations were used on the Perkin Elmer 9600 Thermal Cycler in hopes that optimal conditions for Astacin gene replication would be found; however, after many PCRs with different templates and temperatures were tried, no useable PCR products were obtained.
In order to combat the problems with the primers, a new set of primers was designed using the computer program Mac Vector. They were also based on the Astacus astacus Astacin sequence.
After experimenting with different PCR annealing temperatures for the new primers, it was found that a combination of one of the old primers and one of the newly designed primers gave the most consistent products.
Another aspect of Astacin examined in this project was the possible multiple copy nature of the gene region. It isn’t uncommon to find multiple copies of nuclear genes in a taxa’s genome. In order to work around this possibility, cloning was used on successful PCR products in order to isolate single copies of the Astacin region. The clones were then used as the template for PCR. The cloning showed that the multiple copy possibility was not a major factor to be concerned with; such a conclusion is based on the fact that multiple clones from one PCR product yielded identical sequences.
The obtained sequences were then analyzed by PAUP using the Distance method, Neighbor-joining. Using PAUP, a phylogram was obtained which met the expected results. The sequences representing the family Astacidae were grouped together and those from the family Cambaridae were also grouped together. In the family Astacidae, the members of genus Astacus were grouped together and those of genus Astropodomobious also were grouped together as expected. The members of family Cambaridae including separate groupings of Orconectes species and Cambaroides species as expected. This follows the accepted evolutionary lineage for crayfish.
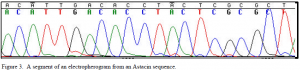
An interesting find from this research was that the published Astacus astacus sequence had an 80 base pair insertion in the intron between exons 4 and 5. The insertion was not present in the sequences obtained in this study. Perhaps the published sequence came from a rare gene copy that was not found in this sequencing or that the insertion was restricted to a specific population of Astacus astacus different from the one used in this study.
References
- Crandall, K.A., Fetzner, J.W., Lawler, S.H., Kinnersley, M., Austin, C.M. (1999). Phylogenetic relationships among the Australian and New Zealand genera of freshwater crayfished (Decapoda:Parastacidae). Australian Journal of Zoology. 43, 199-214.
- Geier, G., Jacob, E., Stocker, W., Zwilling, R. (1997). Genomic Organization of the Zinc-endopepetidase Astacin. Arch. Biochem. Biophys. 337(2), 300-307.
- Swofford, D.L. ‘PAUP* Phylogenetic Analysis using Parsimony and other Methods. (Sinauer Associates: Sunderland, MA.)