J. Daniel Rasband, Department of Zoology
Abstract
Several different methods have been employed in the extraction and amplification of ancient DNA (aDNA). The work in this thesis Is an effort to compare three popular extraction techniques, chelex, guanidine/ silica slurry, and phenol/chloroform, as applied to extracting DNA from the dentin of ancient teeth. Modern and ancient samples were extracted using the three techniques. The extract was then amplified using the polymerase chain reaction (PCR). The data show that of the three extraction procedures, the most favorable results were obtained using the guanidine/silica slurry extraction technique, producing consistently good amplification. The phenol/chloroform extraction was mildly successful, showing amplification in one third to one half of the samples. The chelex extraction showed consistent amplification of the positive control only.
Extraction Procedures
Chelex
A static charge was applied to a sterile pipette tip, which was then placed in a microcentrlfuge tube containing the dentin powder. The powder that would then cling to the tip was placed in 10% chelex suspension (see appendix 1). The samples were vortexed liberally, and were Incubated at 57° C overnight. Following incubation, the samples were again thoroughly vortexed, and were placed in a water bath at 948 C for fifteen minutes. The samples were then spun in a microcentrifuge at 2500 rpm for three minutes. The supernatant was transferred (containing the DNA) to a sterile I. 7 ml microcentrlfuge tube, and was stored at 4° C. Positive controls consisted of two human hair root bulbs added to a I 0% chelex suspension tube. Negative controls consisted of 10% chelex suspension tubes to which nothing was added.
Guanidine/Silica Slurry
The clinging dentin powder was added to I. 7 ml tubes containing 200 f.lol 4M guanidine thiocyanate digestion buffer. The tubes were vortexed and incubated at 57° overnight. Following incubation, 1100 fLl sodium iodide was added to each sample, along with 25 ml silica slurry. Each tube was vortexed for thirty seconds, and was then incubated on Ice for 15 minutes. The samples were then centrifuged at 2250 rpm for two minutes, and the supernatant was discarded. One ml rinse wash was added to each tube, and the tube was vortexed just enough to break up the silica pellet. The samples were again centrifuged at 2250 rpm for 10 seconds, and the supernatant was discarded. The rinsing was continued until the supernatant was colorless. A final wash of 1.5 ml was added and poured off, and the samples were then vacuum dried. The silica pellets were resuspended in 150 fLl HPLC grade water by vortexing, and were incubated at 55°C for thirty minutes to allow the DNA to elute off the silica particles. The tubes were centrifuged a final time at 2250 rpm for three minutes, and the supernatant (containing the DNA) was transferred to sterile I. 7 ml microcentrifuge tubes and stored at 4° C. Positive controls consisted of two human hair root bulbs added to I. 7 ml tubes containing 200 fLl digestion buffer. Negative controls were tubes containing only 200 f.lol digestion buffer.
Phenol/Chloroform/Isoamyl Alcohol Clinging powder was introduced into 1. 7 ml microcentrifuge tubes containing 200 fLI proteinase K digestion buffer. The tubes were vortexed and incubated overnight at 57° C. After incubation, 200 fLI phenol/chloroform/ isoamyl alcohol (25:24: 1) was added to each sample and mixed well. The samples were centrifuged at 2500 rpm for ten minutes, and the organic layer was removed and discarded. This was repeated a second time, with the difference that 150 fLI of the supernatant was transferred to sterile 1. 7 ml microcentrifuge tubes. Forty fLI 3M sodium acetate was added to each sample, followed by 800 fLI 100% ethanol. The samples were then incubated at -20° C overnight. The tubes were centrifuged at 2500 rpm for ten minutes, and the supernatant was carefully poured off and discarded. 400 fLI 70% ethanol was then added to each tube, and the tubes were again spun at 2500 rpm for ten minutes. The rinse was discarded, and the tubes were vacuum dried. The DNA was dissolved in 150 fLI water, allowing one hour for complete dissolution. The samples were then stored at 4° C. Positive controls consisted of two human hair root bulbs added to 1. 7 ml tubes containing 200 j.Ll digestion buffer. Negative controls were tubes containing only 200 fLI digestion buffer.
Conclusion
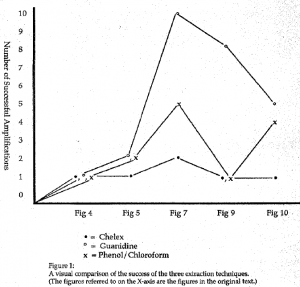
All three extraction techniques that have been used in this study have been used in previous research with positive results. However, it is certain that they have also been applied and negative results were returned. Only the positive results are published. In this study, positive results were also observed with all three extraction techniques. The relative success of the three, however, was dramatically different (figure 11).
Of the three extraction techniques, this study shows the guanidine/silica slurry extraction to be the most effective in extracting and purifying ADNA from dentin powder. The phenol/chloroform/ isoamyl alcohol extraction with an ethanol precipitation and purification was the next best. The chelex extraction did not show significant success in extracting the ADNA from the dentin powder.
Because the three different techniques have returned positive results in previous work, it can only be assumed that the conditions in which the procedures are used play a significant role in the success of the extractions. Also, it should be pointed out that because of the highly sensitive nature of the ADNA and the extremely small quantities at which it is present, even if ADNA is present it does not always amplify (compare the fourth lanes of figures 4 and 8).
In conclusion, when using dentin powder from ancient teeth as a source of DNA, the best, most reproducible results are obtained using the guanidine/ silica slurry method of extraction and purification.
(These pages are excerpts from a thesis by the same title submitted to BYU’s Honors department, a copy of which can be found in the HBLL library.)