Paul B. Savage and Dr. Bryce A. Harrison, Chemistry and Biochemistry
Gram-negative bacteria contain a toxin known as lipid A (1 in Figure 1). During bacterial infections, lipid A may be released from the bacteria and cause a condition known as septic shock, an over-response of the host immune system that may result in death (2). In fact, septic shock kills over 100,000 people annually in the U.S.1 In hopes of developing treatments for septic shock, we are researching compounds that bind and detoxify lipid A. One such compound, polymyxin B (2 in Figure 2), is itself too toxic for medicinal use; nevertheless, we want to understand how polymyxin B binds lipid A. This understanding will help us develop compounds that bind lipid A and can be used medicinally to prevent sepsis. In fact, members of our lab group have already made mimics of polymyxin B that may bind lipid A. We want to test the lipid A binding ability of these compounds.
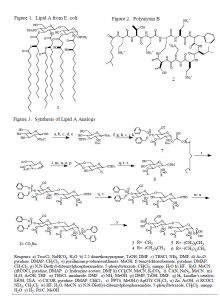
To achieve these ends, we are synthesizing analogs of lipid A (3-6 in Figure 3). These compounds conserve the saccharide head group of lipid A but have shorter hydrophobic tails of varying length. Consequently, the analogs are more water soluble than lipid A and can be used in binding studies where lipid A cannot be used because of its insolubility. Once the analogs are made, we will measure the binding of the analogs and polymyxin B using microtitration calorimetry. Comparing the binding of the analogs with varying tail length will show the role of the hydrophobic portion of lipid A in the binding with polymyxin B. We expect that the binding occurs mostly through ionic and hydrophobic interactions of polymyxin B with the lipid A saccharide head group. Moreover, we will compare the binding of polymyxin B with the analogs in an aggregated and unaggregated state to determine the effects of lipid A aggregation on the binding. Finally, we will assess the lipid A binding ability of the polymyxin B mimics by measuring the binding of the mimics with the lipid A analogs.
For the last two years, we have been developing the synthesis of the lipid A analogs (see Figure 3). This year we extensively revised our synthesis after discovering that reaction c gives a different product than we thought– the silation occurs at the 3 hydroxyl instead of the expected anomeric hydroxyl. Consequently, we had to include additional steps in the synthesis of the glycosyl donor (7 in Figure 3) and completely redo the synthesis of the glycosyl acceptor (8 in Figure 3). However, we overcame these obstacles and are working out the last steps of the synthesis. We have completed up to step w and have attempted step x. Soon we will complete the synthesis and proceed to the testing phase of the project (2).
References
- Schletter, J. et al. Arch. Microbiol. 1995, 164, 383.
- The work of Ryan Marshall, Adam Peters, and Ben Gordon is gratefully acknowledged.