Jonathan Dewey and Dr. Allen R. Buskirk, Chemistry and Biochemistry
Analyzing the function of newly discovered genes and the proteins that they encode is crucial to the advancement of genetics, cell and molecular biology, and medicine. Scientists have implemented various methods to study individual proteins within a cell, including genetic and small-molecule dependent approaches. Our goal is to combine the generality of genetic manipulation with the time-sensitivity of small-molecule dependence to create tools to study the function of Hedgehog pathway proteins and validate them as drug targets.
The Hedgehog (Hh) pathway is involved with patterning and growth in embryonic development and has been shown to be required for the maintenance of some types of adult stem cells, particularly in neural tissues, skin, and blood. Mutations in the Hh pathway allow proliferation of cancer stem cells. The over activation of the Hh pathway is a major factor in various forms of cancer, including that of the stomach, pancreas, and prostate (1). Over activation occurs in ~25% of fatal cancers in humans (2).
Recently, researchers have developed a method to study protein function in vivo using peptide aptamers – inert protein scaffolds that can be evolved to form new protein-protein interactions with a protein target of interest (3). These “peptide aptamers” can be expressed in vivo to bind and inhibit the function of the protein target. We have further developed aptamer technology to add small-molecule regulation.
Murray and co-workers identified peptide aptamers that inhibit the mating pathway of yeast using an inert staphylococcal nuclease scaffold (4). By fusing Murray’s aptamers to an estrogen receptor (ER) we have created peptide aptamers whose function can be controlled by the introduction of estrogen. In an estrogen-free environment, the Hsp90 complex, naturally present in cells, binds to the ER and prevents the aptamer from binding to its target. In the presence of estrogen the Hsp90 complex is displaced and the peptide aptamer is free to bind and inhibit its target (5). In a proof of principle experiment, we took previously characterized peptide aptamers that bind to Sfh1, a protein involved in the mating pathway of yeast, and fused them to the ER. Our preliminary results demonstrated that peptide aptamers could indeed be rendered ligand-dependent. In the presence of alpha-factor, which induces the mating pathway response, colonies formed only when the aptamer blocked signaling. The aptamer was, in turn, only functional in the presence of estrogen.
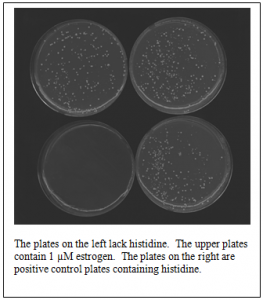
The application of this new technique to the study of the Hh pathway will initially require the performance of a yeast two-hybrid selection to identify a peptide aptamer that interacts with one of the Hh pathway components. The yeast two-hybrid system requires the construction of hybrid proteins that interact with each other to trigger the transcription of a reporter gene. To test the ligand-dependence of the yeast two-hybrid assay, we fused the gene encoding the Sfh1 component of the yeast mating pathway to a DNA fragment encoding the DNA-binding domain of Gal4. We also fused our estrogen-dependent, Sfh1-binding aptamer to a DNA fragment encoding an activation domain. Interaction between Sfh1 and our aptamer would activate HIS3, a reporter gene that allows cells to grow on plates lacking histidine. We transformed yeast cells with both vectors and plated the cells on media lacking histidine. Half of the cells were plated on media containing 1 µM estrogen while the other half were plated on media lacking estrogen. As anticipated, colonies formed on the plates containing estrogen, but no colonies formed on the plates lacking estrogen (see figure).
In order to identify a peptide aptamer that will interact with a component of the Hh pathway, we will perform similar two-hybrid selections. We have created new DNA-binding complexes by replacing Sfh1 with Suppressor of Fused (Sufu) and Gli, two components of the Hh pathway. The activation domain will be fused to a new peptide aptamer library, consisting of a randomized peptide insert in the active site of the protein scaffold. We will prepare the random peptide insert by self-annealing the following oligonucleotide: 5’-CCC GATTCTTCGGTGGT(NNS)16GGTGGTGTCGACACC-3’ [N = A:T:G:C (1:1:1:1), S = G:C (1:1)]. Annealed DNA will be extended using the Klenow fragment of DNA polymerase I. The double-stranded product will be cut with Eco RI and Sal I. The library will be ligated into the staphylococcal nuclease construct (pNER-AD) cut with Eco RI and Sal I (4). Any cell containing an interacting combination of a Hh component and a peptide aptamer will activate HIS3 and allow cells to grow on plates lacking histidine.
The development of small molecule-dependent peptide aptamers will facilitate the study of the Hedgehog pathway, as well as other protein signaling pathways. Using our peptide aptamer, scientists will be able to disable and reactivate the Hedgehog pathway at a desired time point through the introduction of estrogen. The generality of protein-protein interactions will enable researchers to easily create and identify peptide aptamers specific to other cell-signaling components. Ligand-dependence will then allow researchers to study a specific signaling pathway in a time-sensitive manner.
