Bryce A. Harrison and Dr. Paul B. Savage, Chemistry and Biochemistry
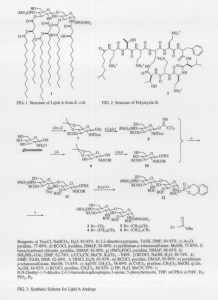
Lipid A (1 in Fig. 1) is a bacterial compound that can cause sepsis, an overreaction of the immune system during bacterial infections.1 Because no good treatment is known, sepsis causes over 100,000 deaths annually in the U.S.1 Treatments for sepsis are difficult to develop partly due to a lack of understanding of how lipid A interacts with other molecules. Consequently, we are synthesizing lipid A analogs (3-6 in Fig. 3) to studying the mechanism of binding of lipid A to polymyxin B (PMB, 2 in Fig. 2), a naturally occurring cyclic peptide. Understanding the mechanism of binding will aid in developing medicinal compounds that bind and detoxify lipid A. In addition, PMB shares many structural similarities with the active sites of immune system proteins that bind lipid A.2 Thus the lipid A-PMB system may serve as a model for understanding immune system interactions with lipid A.
The binding of lipid A and PMB is difficult to study directly due to the low solubility of lipid A in aqueous solutions. At concentrations high enough to measure binding constants, lipid A aggregates, making unclear the form of lipid A involved in binding. To overcome this problem, we want to perform studies using lipid A analogs which are more soluble but exhibit the same mechanism of binding. The binding occurs principally through the saccharide head portion. Consequently, the analogs conserve the saccharide head group. However, to increase solubility, the analogs have a reduced, unbranched hydrophobic portion of variable length. Once the analogs are synthesized, we will measure the binding constants of the analogs with PMB by microtitration calorimetry. We will be able to verify the minimal effect of hydrophobic chain length on binding by comparing the binding of the different analogs. In addition, we plan to study the effects of aggregation of lipid A on the binding by comparing the binding of the analogs above and below the concentration at which they aggregate.
In preparations for the studies, we have achieved a functioning scheme for the synthesis of the lipid A analogs (see Fig. 3). Beginning with glucosamine, we functionalize and protect each half of the molecule. We then couple the two halves, functionalize the remaining positions, and deprotect to give the desired product. To date we have synthesized through compound 12 (in Fig. 3) in the synthesis of 5 and are well advanced in the synthesis of all of the analogs. Moreover, we can use similar schemes to synthesize lipid A and monophosphoryl lipid A analogs for further studies.3
References
- Schletter, J. et al. Arch. Microbiol. 1995, 164, 383.
- Little, R. G. et al. J BioL Chem. 1994, 269, 1865.
- The aid of Sanjoy Mullick, William Thomas, and Adam Peters is gratefully acknowledged.