Christensen, Zachary
An Open Source Function Utilizing Random Field Theory for MRI Analysis
Faculty Mentor: Erin D Bigler, Psychology Department
Introduction
When looking at an MRI scan of the brain one is actually viewing millions of voxels
(three dimensional pixels) that represent individual groups of signals produced by the
MRI machine. Therefore, each voxel is representative of the brains composition at that
point in three dimensional space or metabolic activity (as is the case in functional MRI).
These qualities are utilized in research to give anatomically meaningful comparisons of
the brain between individual test subjects. Doing so involves the process of mass
univariate testing and produces a single statistical value at each voxel representing the
univariate test (i.e. a tvalue). However, such an approach to comparing MRI scans
creates an issue of multiple comparisons and interpretability.
Results from scientific studies are often reported in terms of the probability (pvalue)
that the results are just due to chance. For example, a common statistical threshold
used in the social sciences is a pvalue of .05 and when used is equivalent to reporting
that the results are deemed significant because there is only a five percent chance that
the results are due to chance. However, with each additional analysis the probability of
producing a false positive increases. There are several approaches to correcting for
multiple comparisons, such as dividing the pvalue threshold by the number of tests
(Bonferroni correction). Applying a correction method such as Bonferroni to mass
univariate testing in MRI scans would result in a threshold of less than 5 x 10^8.
Not only is such a threshold unrealistically low, but this method of correction treats each
voxel as a completely independent variable with no relation to its surrounding voxels.
Additionally, even if there were no issue of multiple comparisons interpreting millions of
individual statistical tests in such a way that produces anatomically meaningful
information is impossible without some clear heuristics.
Random field theory (RFT) allows the “field” of statistical values resulting from mass
univariate testing to be treated as random noise containing potentially significant signals
within it. Such an approach accounts for the spatial properties of mass univariate testing
and provides a foundation upon which increased interpretability of outcomes can be
made. Despite the widespread use of mass univariate testing in MRI research, RFT
techniques are often only available to those able to pay for the use of expensive
software. Such limitations reduce the amount of reasonably reproducible brain MRI
research.
In order to remedy these issues we provided additional functionality within the
open-source package ANTsR that extends R’s existing statistical framework into RFT
based analyses for brain MRI scans. The following demonstrate a basic RFT analysis
utilizing this newfound functionality.
Methods
Data was obtained from the pediatric template of brain perfusion (PTBP; Avants et al.,
2015). All participants in the PTBP were screened for neurological and psychiatric
diagnoses and underwent acquisition of a structural MRI scan using a Siemens 3T TIM
Trio scanner (818 years of age). The Advanced Normalization Tools (ANTs; Avants et
al., 2011) software was used to prepare images for statistical analysis.
Images were imported into R using the ANTsR package. A simple linear model was
used to find the correlation between each participant’s MRI scan and age. Familywise
error correction was used to correct for multiple comparisons via implementation of RFT
as previously discussed. The code used to implement the analysis is available in
Appendix A.
Results
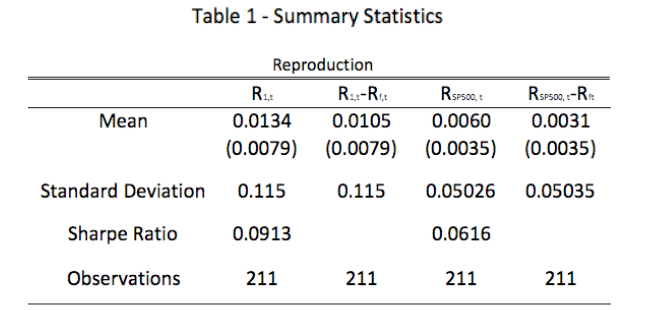
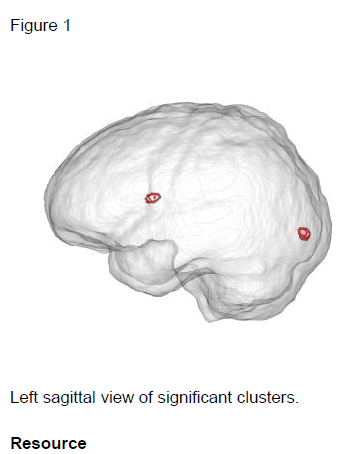
Two statistically significant clusters were found proximal to the occipital pole and
anterior corpus callosum (see Table. 1 and Figure 1). Each cluster represents
increased anatomical deformation correlated with age.
Conclusions
These findings validate previous studies that demonstrate neuroanatomical changes to
be associated with age (Kandel et al., 2013). More importantly, this replication supports
the validity of the statistical analysis being used. The 1,000+ lines of code contributed
through this study has resulted in opensource software that allows anyone to access
and scrutinize its development. This increased accessibility should result in faster
improvements to computational neuroimaging and decrease the financial burden of
performing such analyses.
library(ANTsR)
ptbp <read.
csv(file.path(root, “data/demographics.csv”), header = TRUE)
ipaths <paste(
root, “rft/warp/”, ptbp$SubID, “_”, ptbp$ScanDate, “.nii”, sep = “”)
mask <antsImageRead(
ipaths[1])
mask[mask != 0] <1
imat <imagesToMatrix(
ipaths, mask)
wb_bool <rep(
TRUE, nrow(imat)
mydata <mydata
<imgDataMake(”
wb”, imat, mask, wb_bool, ptbp)
fit % rftLm(wb ~ AgeAtScan)
contrastMatrix <matrix(
c(0, 1), nrow = 1)
myresults <summary(
fit, contrastMatrix, cthresh = 100)
Resources
Avants B. B., Duda J. T., Kilroy E., Krasileva K., Jann K., et al. The pediatric template of brain perfusion. Sci Data. 2015;2:150003. PubMed PMID: 25977810; PubMed Central PMCID: PMC4413243.
Avants B. B., Tustison N. J., Song G., Cook P. A., Klein A., et al. A reproducible evaluation of ANTs similarity metric performance in brain image registration. Neuroimage. 2011 Feb 1;54(3):203344.
PubMed PMID: 20851191; NIHMSID: NIHMS238501; PubMed Central PMCID: PMC3065962.
Kandel B. M., Wolk D. A., Gee J. C., Avants B. Predicting cognitive data from medical images using sparse linear regression. Inf Process Med Imaging. 2013;23:8697.
PubMed PMID: 24683960; NIHMSID: NIHMS725768; PubMed Central PMCID: PMC4603981.