Robert Fuller and Dr. Alonzo Cook, Chemical Engineering
I. Introduction
Decellularized porcine tissue is currently used for allografts of tendons, blood vessels, and dermal tissue. The success of these procedures has been attributed to the conservation of collagen structures across mammalian species, making porcine collagen nearly identical to that of humans. Signaling chemicals for cellular differentiation are as well identical among most mammalian organs. This has led to the idea of partial or whole organ regeneration through a donor collagen matrix.
As over 100,000 persons in the US alone are diagnosed with end-stage renal disease annually, there is a large demand for replacement kidneys that is only slightly reduced by available transplant organs. This project sought to find an improved treatment for whole organ decellularization. Issues arose from our initial project proposal that required significant modification, but results suggesting a potentially improved treatment remain.
II. Methods
Initial experiments were conducted with 0.1 wt% EDTA solutions. These included EDTA with 0.5 wt% SDS (sodium dodecyl sulfate), EDTA with 0.5 wt% SDS with 30 minute hypertonic/hypotonic cycling solutions, and EDTA with 1 vol% Triton X-100, as well as using the EDTA alone. After several trials, all of these solutions showed either no significant improvement with EDTA or failed to approach decellularization after 12 hours. This showed an ineffectiveness of EDTA in concentrations not expected to interact with the collagen matrix. As well, variations in the size of porcine kidneys obtained from a local abattoir also introduced a convoluting variable in these initial experiments.
The project was thereafter modified to be performed on a section of porcine kidney tissue immersed in solution on a vial shaker for 24 hours. The sectioning removed the variable of kidney size, while the treatments were changed to several more comparable with recent literature. The treatments used involved 0.05 wt% EDTA and Trypsin, 0.1N NaOH, 1% vol% peracetic acid (PAA), and 3 vol% Triton X-100.
After the treatment, they were subjected to assessments including (a) H&E staining for determining magnitude of decellularization, (b) SEM imaging, (c) PicoGreen® assay for residual DNA content, (d) Collagen quality stains for 1. Overall collagen quantification, 2. Glycosaminoglycan (GAG) quantification, 3. Elastin content, and finally (e) recellularization with fluorescing human Renal Cortical Tubule Epithelium (RCTE), with fluorescence measured for estimating cell growth. After 5 trials of each quantifying assay, the data were analyzed with JMP and Excel using a 95% confidence interval depicted on all graphs.
III. Results
(a) H&E staining: The peracetic acid treatment was the only one of the five treatments found to not remove all cell clusters, with some cells left behind in the Triton X-100 and Trypsin/EDTA treatment.
(b) SEM imagining: Fibrous tissue was found in the peracetic acid treatment, partially fibrous tissue in Triton X-100 and Trypsin/EDTA treatments, and nonfibrous tissue in NaOH tissue.
(c) PicoGreen® assay: The NaOH treatment was most successful in removing DNA, while Triton X-100 and Trypsin/EDTA treatments had poor DNA removal. Results are shown in Figure 1.
(d) Collagen Quality Stains with quantification performed by extracting dye adsorbed for collagen and GAG content were used to compare effects of chemical treatments.
(1) Sirius Red and Fast Green staining was used to assess collagen content and extracellular proteins. The peracetic acid and Trypsin/EDTA treatments were the only samples with visually detectable amounts of extracellular proteins remaining. Quantified results are depicted in Figure 2.
(2) GAG content was visually misleading, as the peracetic acid appeared to adsorb most of the dye. Quantification showed that the Triton and SDS treatments maintained the most GAGs as shown in Figure 3.
(3) Orcein-Elastin staining gave nearly identical results to GAG assays visually, but was not quantified in these experiments. They showed considerable elastin dye in the peracetic acid and Trypsin/EDTA treatments, but this may be more due to chemical interactions rather than actual elastin content.
(e) Recellularization was measured by relative fluorescence of cells, examined on a daily basis. The NaOH treatment exhibited the greatest growth, followed by the peracetic acid treatment. The other treatments had comparatively poor cell growth, with Triton having slightly improved growth. The results are depicted in Figure 4.
IV. Discussion
The NaOH treatment was highly successful in recellularization, despite its overall poor structural preservation. All other solutions were either acidic or nearly neutral, but the basicity of NaOH may be more effective of removing the weakly acidic DNA and RNA, while leaving weakly-basic protein intact. This treatment is rarely used in current literature, and therefore may be the main subject of future research.
Peracetic acid oxidizes tissue, which caused breakdown of collagen visible in the SEM imaging. The subsequent increased surface area may be the cause for its vivid color in staining, and may affect the results of quantifiable assays. Despite its incomplete decellularization, the mixed effects of peracetic acid warrant further investigation.
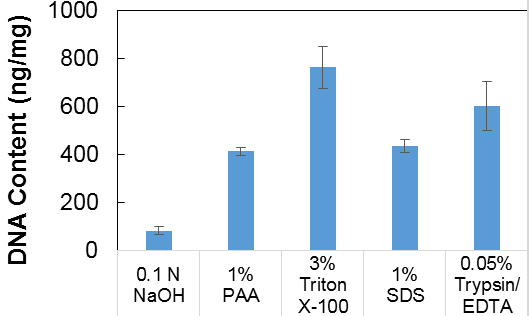
Figure 1: PicoGreen® assay for DNA content of decellularized tissue.
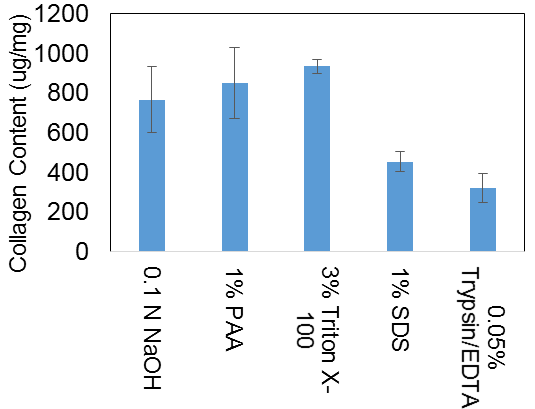
Figure 2: Estimated collagen content of decellularized tissue
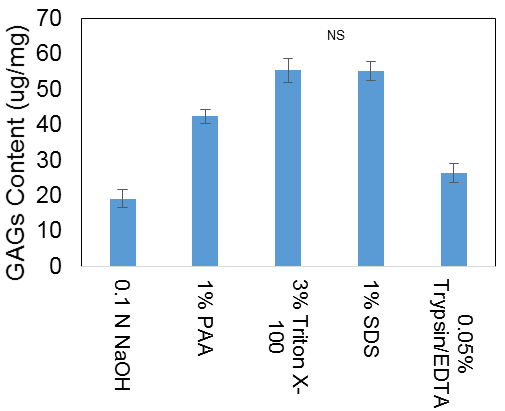
Figure 3: Estimated GAG content of decellularized tissue
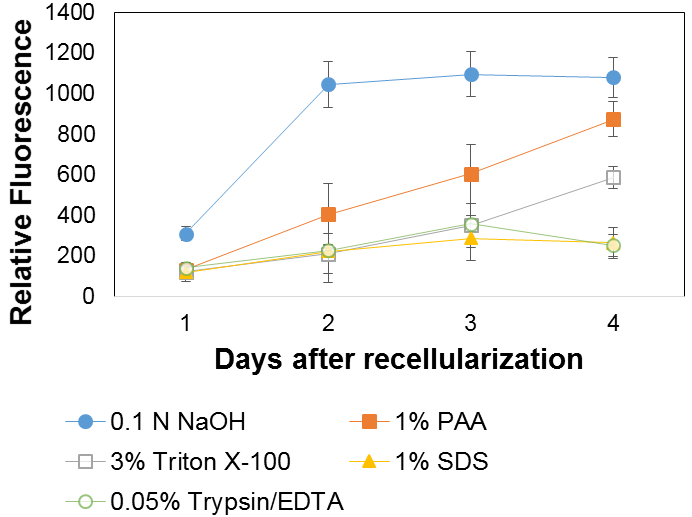
Figure 4: Relative cell fluorescence over time when incubating cells with human renal cells.
V. Conclusions and Acknowledgements
Despite its theoretical significance, EDTA failed to visibly affect decellularization in initial trials with SDS, Triton X-100, and when used alone, each at 0.1 wt%. Considerable evidence was found for NaOH as an effective decellularization agent. Despite its damage to the collagen structure, it showed a superior final effect on recellularization, and will be subject for further study. We would like to thank Nafiseh Poornejad, Evan Buckmiller, Lara Schaumann, and Angella Nakalembe for their guidance and assistance throughout this project.
