Devon Blake and Dr. Barry Willardson, Chemistry and Biochemistry
Introduction and Project Importance
Cells detect and respond to a myriad of extracellular signals via seven-transmembrane G protein-coupled receptors and their associated G protein signaling pathways. The pathway is initiated by the binding of a signaling molecule, such as a hormone or neurotransmitter, to its binding site on the extracellular side of the receptor. This interaction activates the G protein heterotrimer, composed of Gβ, Gβ, and Gβ subunits, which functions by propagating the signal throughout the cell. First, however, the G protein heterotrimer must be assembled from its nascent polypeptides. The initial step in this process is the formation of the Gβγ dimer. Gβγ is an obligate dimer in which neither subunit is able to fold into a stable structure on its own. A fundamental question that has persisted for years is how the Gβγ dimer can form when the individual subunits are unstable. Recently, however, strong biochemical evidence from our lab and others showed that the Gβγ-binding protein, phosducin-like protein 1 (PhLP1) acts as a co-chaperone with the cytosolic chaperonin CCT to fold Gβ and promote its interaction with Gγ (1). Since then, we have spent considerable time and effort to uncover the molecular mechanism by which PhLP1 and CCT work together to assemble Gβγ dimers. We had determined the structures of two intermediates in the Gβγ assembly process, the Gβ-CCT complex and the PhLP1-Gβ-CCT complex, by cryo-electron microscopy to 13 Å resolution (3), but even with this major achievement the exact binding site of Gβ remained a mystery.
Methods
Chemical cross-linking combined with mass spectrometric analysis is an attractive technique for obtaining structural information on proteins and protein complexes (4). Cross-linked proteins can be enzymatically digested, and the cross-linked peptides obtained can be analyzed by mass spectrometry to identify both the crosslinked peptides and the site of cross-linking. To solve the exact binding site of Gβ, we used this type of crosslinking. We started by synthesizing cDNA constructs for the genetically engineered proteins with appropriate affinity tags: CCTα with a calmodulin-binding protein (CBP) tag, PhLP1 with a c-terminal Myc-TEV-HIS6 tag and Gβ with a c-terminal HPC4 tag. The cDNAs were then inserted into a Bac-to-Bac® Baculovirus Expression System (Invitrogen), and Hi-5 insect cells were transfected with the recombinant bacmid DNA to generate recombinant protein. After lysis of the cells, the lysate was submitted to tandem affinity purification using a calmodulin affinity column to isolate CCTα-containing complexes, followed by a Co2+-chelate column for HIS6-tagged proteins to isolate the subset of complexes with both CCTα and PhLP1, and finally an anti-HPC4 antibody column for HPC4-tagged Gβ to isolate only those complexes containing CCTα, PhLP1 and Gβ. Following purification of the complex, it was be reacted with the cross-linking reagent CBDPS-H8/D8 (Creative Molecules, Inc.). After this reaction, the complex was digested with the protease trypsin, and then subjected to affinity chromatography on a streptavidin column. The cross-linked peptides, which contain a biotin moiety on the cross-linking reagent, were retained on the column through the high-affinity interaction of biotin with streptavidin while the non-cross-linked peptides passed through the column. The cross-linked peptides were eluted with excess free biotin and analyzed using mass spectrometry and subsequent computational analysis to identify the cross-linked residues.
Results
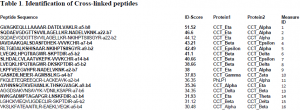
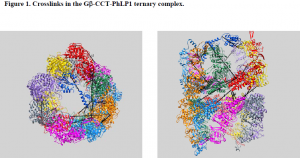
We were able to express all three proteins concomitantly in insect cells and were able to isolate sufficient ternary complex to perform mass spectrometry analysis. However, the procedure was not as straightforward as it could have been. We ran into several problems which we had to troubleshoot. One problem was the buffer that we used to store the complex. When we tried to analyze the complex using mass spectrometry, the buffer formed a polymer that occluded the signals from our peptides of interest. It took a while to troubleshoot the problem, but once we realized that it was the buffer we were able to proceed using a different buffer system. Another big problem was that the mass spectrometer was not functioning for a few months. Thus, we had to take our prepared sample up to the lab of Dr. Sarah Franklin at the University of Utah for mass spectrometry analysis. Despite the setbacks we encountered, we have been able to identify several important crosslinks (Table 1), which we have fit into the structure of Gβ-CCT-PhLP1 we obtained from docking their crystal structures into the cryo-electron envelope (Figure 1). Before our data is published, we need to identify a few more crosslinks. Then, it will be submitted to the journal Molecular Cell.
Conclusion
In order to fully understand the mechanism by which Gβfolds into its native state, the exact binding site inside the CCT complex must be known. Given the importance of chaperone-mediated protein folding and complex assembly as well as the vital role of Gβγ dimers in G protein signaling, this study is very important as this folding pathway could serve as a potential target for future pharmacological studies.