Tim Rand and Dr. Barry Willardson, Chemistry & Biochemistry
Introduction
The theory of evolution by natural selection is arguably the most important scientific theory ever formulated. Its importance has not been manifest in an “ooh, ah” technological way—it doesn’t empower us to fly airplanes or to illuminate streets at night—but no scientist really thinks technology is the sine qua non of science anyway. The really big theories are the ones that change our thinking, rather than the way we get around or see at night. The theory of evolution changed the way we think about life itself. Interestingly, our acceptance of evolution is based mainly on its elegance. As far as evidence goes, it is circumstantial—although available by the truckload. The challenge of finding more direct evidence is multifaceted. Until the 1944 discovery that DNA was responsible for heredity, no one knew where to look for more direct evidence. There is also a problem inherent to archeology in general, but amplified in the case of molecular archeology. Namely, how do we show what happened in the past given only random shards existing in the present? Finally, there are some rather difficult scientific questions: when species B evolves from A, what exactly changes in the process? Does it make different proteins that cause a corresponding morphological or behavioral difference? Or are the differences in sequence insubstantial, while the important differences are in the regulation of the cognate proteins (i.e. when they are produced and degraded)? Solving these issues and finding molecular evidence of evolution is left as the Sysiphean (1) task for modern evolutionists.
In 1972, Susumu Ohno published a book describing how new proteins might evolve by gene duplication. The mechanism is important to our understanding not only of how a species might evolve, but the related problem of how more complex biological systems might evolve from simpler ones. His theory states that proteins with new functions cannot evolve until a gene is duplicated. The reason is simply that changing a useful protein to something different will destroy the original function and therefore sacrifice the fitness of the organism. However, if the gene encoding the protein is duplicated, one copy can retain the original function and the other can act as a pseudogene—randomly taking on mutations with the potential of gaining a novel function beneficial to the organism. At the time, Ohno was only able to offer elementary examples for support of this phenomenon. The main example used throughout the book was the globin family (hemoglobin). Unfortunately, while these genes share sequence similarity suggesting a common origin, there is not a definitive difference in function—all encode oxygen-carrying molecules that differ only slightly in their affinity for oxygen.
Phosducins
Recently, two proteins were discovered which might provide better evidence of protein evolution occurring through gene duplication. Phosducin (Pd) is a regulator of the vision pathway. The biological role of its homologue, Phosducin-like protein (PhLP), is not yet understood. However, a sequence alignment provides us with clues as to the functions that have and have not been preserved between both products of this putative gene duplication (see Figure 1).
According to the work done on phosducin, we know that its amino terminus, especially a helix one, is important for its ability to bind to the bg subunits of heterotrimeric G proteins. Binding to Gbg by phosducin inhibits Gbg-dependent turnover of the a subunit and the light signal passed through the activated a subunit (see The Cell 737-739 for review of G-protein signaling). Its carboxy-terminus also seems to play a role in Pd’s function. It forms a large protein glob (similar in structure to thioredoxin) that may act in de-localization of Gbg from the cytosolic surface of the membrane. The sequences of these two components, a helix one and the carboxy-terminus, are similar when compared via alignment of PhLP and Pd (see Figure 1).
The fact that the a helix (red arrow Figure 1) remains identical between the two proteins suggests a strong selective pressure that has maintained the bg binding function in both proteins after divergence. The semi-conservation of the C-terminus (see green arrow in Figure 1) offers more cryptic information. Within this region considerable differences have accumulated between PhLP and Pd sequences, and yet there remains enough similarity to positively identify the two as cognates. If this domain acts merely as a protein blob to push the bg subunit off the membrane (as hypothesized above), it may have a low number of functionally constrained sites. In other words, many of the amino acids could mutate without deleteriously effecting the overall fold or the function of that domain. One of the members of our lab is now investigating the function of the N-terminal and C-terminal domains in Pd and PhLP using an in vitro experiment. His results may provide circumstantial evidence to support this theory. More direct evidence might also be provided by the determination of PhLP·Gbg structure via X-ray crystallography.
A Molecular Clock?
If the mutations have indeed mostly occurred in functionally unconstrained positions leaving the overall folding intact and therefore allowing neither positive or negative selection to occur, we are justified in using the mutations in this domain as a molecular clock. Linus Pauling and Emile Zuckerkandle proposed the existence of molecular clocks in 1962, and though certain limitations apply, many proteins and DNA sequences have since been shown to accrue mutations in a clock-like fashion. Molecular clocks have been used to date the divergence of various species, mostly in hopes of clarifying phylogenetic trees otherwise based on partial fossil information. Here, we will use it to date the duplication of the primordial phosducin gene.
A phylogenetic tree was developed (see Figure 2) using the HKY-85 model for DNA mutation and PAUP (phylogentics program). The molecular clock hypothesis was not negated by a P value test and is therefore useful for our dating procedure. All known Pd and PhLP C-terminal DNA sequences were used, but the third position of each codon was ignored because it contains less reliable data in long-term divergence studies. Using the numbers associated with each branch and the putative date of mammalian radiation, more specifically the human-rat divergence of 75 million years ago (MYA), we can calculate that the Pd gene duplication event occurred ~850 MYA. This date should be considered the most ancient date possible as it assumes a constant rate of mutation throughout the history of two genes after duplication. In reality, the duplicated gene most probably acted as a psuedogene during the early portion of its existence. This suggests that some fraction of the mutations would have occurred abnormally rapidly at first and caused an over-prediction of the actual date using this approach.
Phosducin Fishing And Future Research
Using Pd specific and PhLP specific antibodies, representative species from various classes of animals were checked for Pd and PhLP production. The most ancient class of organisms found to produce Pd and PhLP was fish (which diverged from the tetrapod lineage 450 million years ago). Interestingly, Pd was not present in amphibians, while PhLP was. Finally, PhLP was found to be the more ubiquitously expressed protein, detectable in the majority of rat tissues, while Pd was only found in retina tissue.
The fact that PhLP is more ubiquitously expressed argues that its role in the physiology is more basic, working as a general G protein signal regulator, while Pd’s role is specific to the visual system. This image of PhLP being more basic to the physiology of vertebrates is supported by the fact that amphibians have maintained expression of PhLP but seem to have lost the Pd gene from their genome altogether. With a wider range of expression it seems safe to assume that PhLP had its biological role determined early on and that its function may closely resemble the original pre-duplication gene. What is its physiological importance? If the two proteins were expressed inside the same cell, would they ever have opposite, or at least distinguishable, effects?
We have checked PhLP for phosphorylation by protein kinase A (PKA), the regulatory kinase of Pd. The results showed that PhLP is phosphorylated at a rate about 1/5 as fast as Pd. However, its phosphorylation is not inhibited in vivo using PKA inhibitors. This suggests that the two proteins do in fact play out distinguishable roles in the physiology of an animal—Pd is PKA sensitive while PhLP is not.
We hope to verify the duplication date analysis by adding more sequences to allow for statistical verification of constant mutation rates between species in these two genes. We also are continuing work toward the determination of functional differences between Pd and PhLP. We also are sequencing fish Pd as it would be of great interest not only for these studies, but also to those doing research in vision physiology as a majority of such researchers are using fish for their model.
Complex Systems
For many people a strong counter-argument against evolution is the existence of multi-component complex systems. The visual signal cascade in mammals is such a complex system involving rhodopsin, transducin, phosphodiesterase, cation channels, and phosducin (along with other players including GAP, Na+/Ca+ exchannger, arrestin, rho kinase and some perhaps not yet discovered). Developing such a complex cascade requires small yet beneficial changes which, over time, result in the complete system. Still, there is no obvious small beneficial change that can be made to the genome of an animal without the components for at least a partial visual system already present. For instance, making the primeval-rhodopsin a little more like the modern version of rhodopsin doesn’t do any good unless transducin is already there, but transducin couldn’t have evolved without the selective advantage provided when it co-exists with rhodopsin (and the other components too). “Therefore, from nothing can come nothing,” is argued.
In fact there is a critical flaw in the assumptions of this argument: that there is no selective advantage to having one of the components without having the rest. We must always remind ourselves that the presently understood role of a protein is not necessarily the only role it has or has had. If each of the components of a complex system had, at some earlier time, another role distinct from that of the present complex system, we can avoid the paradox altogether. For instance, rhodopsin, or a rhodopsin-like molecule, might have been selected without any of the other visual pathway molecules present (Nature 1997 Jun 2; 387(6634):667-668 Rhodopsin evolution in the dark. Crandall KA, Hillis DM). Our research supports the possibility of a similar process resulting in the creation of the components of complex systems by emphasizing the role of gene duplications. We hypothesize that phosducin-like protein filled a rather generic molecular niche in a wide array of cell types and that phosducin was created by gene duplication followed by a period of existence as a psuedogene resulting in the gain of function making it specifically useful as a visual system component. Subsequent natural selection played a role only in fine-tuning that function (mostly along the Nterminus) resulting in the present day phosducin. Such a process could produce improved systems or even produce entirely new systems by continually mixing and matching new proteins made from pre-existing ones.
Thanks to Barry Willardson and Keith Crandall for invaluable help with various technical aspects of this research. I am also indebted to ORCA for the funding that made this work possible and the opportunity to share the results with others.
References
1. Sisyphus was a Greek mythological character who was given the arduous task of rolling a boulder up hill as a punishment for attempting to trick the gods. “Sisyphean” is referring to such an endlessly difficult task
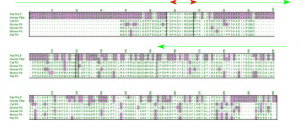
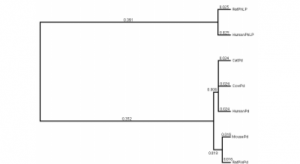
FIGURE 1. PhLP and Pd amino acid sequence alignment. The top two sequences are PhLP and the bottoms five are Pd. Residues highlighted in purple are the minority at that position (putative mutations). The only exceptions are positions where there are three of one amino acid and four of another, in which case the amino acid represented three times is considered the original amino acid if it was found in both the PhLP and Pd sequence (see position # 238). The red arrow signifies a helix one, which is important for Gbg binging. The green arrow signifies the C terminal domain where considerable amino acid sequences conservation is observed (uncolored positions). Residue 120 (ser 73 of Pd) is boxed to denote an important difference that provides the two proteins differentiated regulatory mechanisms.
Figure 2. PhLP/Pd Phylogenetic Tree. All numbers (representing relative mutation) along branches from RatPin Pd to Human Pd are added (0.016+0.018+0.009+0.024=0.067) to give a rate of mutation since human/rat divergence (0.067/75MYA=0.00089/MY). The numbers from Human Pd to Human PhLP are then added (0.025+0.361+0.352+0.009+0.024=0.771) and this number is divided by the rate of mutation (0.00089/MYA) to give an approximate date of 863 MYA for the Pd PhLP gene duplication. This date should be considered the most ancient date possible because it assumes constant mutation throughout time though it is likely one of the genes mutated more rapidly
