Jason Burton and Professor Marc Hansen, Department of Physiology and Developmental Biology
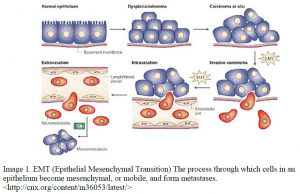
Cancer is one of the leading causes of death in the United States. This is mostly due to a lack of understanding into the function of cancer and the different mechanisms it uses to spread. The process by which cancer cells detach from the primary tumor and spread to form metastases is known as Epithelial Mesenchymal Transition (EMT). EMT is a process whereby specialized epithelial cells undergo a transformation back to primitive, mobile cells (see figure 1). These cells can travel through the basement membranes and enter the nervous and vascular systems and spread throughout the body. This process has been well documented in MDCK (kidney) cells and can be induced by stimulating the cMET pathway by HGF (Hepatocyte Growth Factor) treatment; however, a comprehensive picture of gene activity and splicing events during EMT has not yet been compiled.
My proposed research was to design an experiment that would analyze the gene expression through various time-points of EMT. This would allow us to form a more complete picture of the processes that lead to detachment and invasion of cancerous cells into neighboring tissues. Understanding the changes in gene expression could lead to development of new targets for drug therapy and a reduction in remission of cancer. To compare the gene expression of MDCK cells undergoing EMT, we will extract mRNA from cells that have been treated with HGF for varying amounts of time, convert the mRNA to cDNA, and then sequence the DNA using Illumina Next Generation technology.
To harvest the mRNA from cells, the cells must be scraped off of the plates and spun in a micro-centrifuge to pellet the cells at the bottom of the snap-cap. Cells are then lysed with detergent and treated with RNAse to preserve intact mRNA. The cell slurry is then run through the Invitrogen RNeasy kit and isolates total RNA. We then use the Illumina Tru-Seq kit to isolate mRNA from that sample and convert the samples from single-stranded RNA to double-stranded cDNA. These samples have tags attached to the end for identification after sequencing. Samples are pooled together and mailed to Boston University where the Sequencing Core Lab sequences the DNA and returns the data. Each run of the Illumina Sequencer produces around 500 GBs of data and will need to be interpreted to define what interactions are occurring. We’ll be looking for up and down-regulation of key genes involved in cell motility and function, as well as different splice variations of RNA.
Unfortunately, the predicted timetable for completing this experiment and acquiring publication quality data was not met. Multiple road blocks were encountered along the way; some were beyond my control and others could have been fixed with better planning.
Whole mRNA sequencing techniques are brand new to the Hansen lab, and mistakes are always inherent with a first attempt in anything. An early mistake that occurred was in ordering the wrong sequencing prep kit. Illumina makes multiple kits that are used in mRNA sequencing experiments, and through lack of research and an acquisition of increased knowledge at a later point, we realized we had ordered the wrong kit. Time was used to try and make the kit work, but when it was determined it would be best to replace the kit, we decided to do so. This was followed by successive contaminations of cell growth media and the incubator, which delayed our ability to grow and prepare cell samples by two or three months. Once we had the correct kit, all the reagents needed and cells prepared, the mRNA isolation and cDNA preparation went according to plan, though delayed. These samples were mailed to Boston University at the end of May where a collaborator is working with us to run the samples through the sequencer. After waiting in the queue for a couple of weeks, the samples were run through the machine, only to encounter more complications and delays.
The Illumina machine has an attached hard drive and computer processor that converts raw data into a usable format. Data must then be extracted from that hard drive and run through another computer program before it is available to view and analyze. The hard drive associated with the machine failed and raw data had to be stored on a separate hard drive until the first was repaired. This caused a back-up in samples that needed to be run and in data that was being exported from the lab. Once that hard drive was repaired, and the data was fed through the processor, there was another fail in the secondary computer program. This means that the data was available but still not on a usable format.
Due to these delays and problems, the data is currently at this state; available, but unusable. We expect to receive the data any day now and will begin data analysis. This will be time consuming, requiring many hours of work on the computer. We hope to be able to identify factors that play a significant role in EMT and the spread of cancer to metastatic tumors. Further research will be done to compare those proteins to known pathways and determine the next steps from there. Our findings will be published in a peer reviewed journal.