Mark Terry and Dr. Michael Whiting, Integrative Biology
Thysanoptera (thrips) is an order of insects currently comprised of 7400 species, 1200 genus-groups, and nine living families with three fossil families (Mound, 2005). Two major suborders are recognized in Thysanoptera: Tubulifera (thrips with abdominal segment 10 tubular in shape) and Terebrantia (thrips with abdominal segment 10 subdivided ventrally). Thrips are commonly regarded as flower-living, though a wide range of biologies have been documented including feeding on leaves, flowers, fungal hyphae, fungal spores, mosses, predators of other small arthropods (Mound and Marullo, 1996) and in one instance, as an ectoparasite of other insects (Izzo et al., 2002). It is well known that the classification of Thysanoptera is generally in a state of chaos (Morris and Mound, 2003), and the monophyly (common ancestry) of the suborders and families have never been formally established. Moreover, phylogenetic (evolutionary development) relationships are virtually unknown, and there has yet to be a formal phylogenetic hypothesis put forth for this group. Given that thrips have such a wide diversity, combined with their unique and highly rearranged mitochondrial genomes, (Shao et al., 2002) a robust phylogeny is essential to better understand the evolution and diversification of this order.
The goal of this work is to investigate the utility of molecular data to resolve relationships within Thysanoptera. Specifically: 1) reconstruct a molecular phylogeny for thrips based on three genes, 2) test for the monophyly of Tubulifera and Terebrantia, and 3) test for the monophyly of families and some subfamilies.
A total of 37 taxa were selected from 10 families representing both suborders Tubulifera and Terebrantia. Six of the nine recognized families were included in this analysis. The tree was rooted to Hemiptera and Psocoptera which are related insect groups.
DNA sequence data were generated for three genes: 18S rDNA, 28S rDNA, and Histone 3. DNA was extracted using the Qiagen Dneasy extraction kit. Using Polymerase Chain Reaction (PCR) diagnosis, the DNA fragments were amplified in thrips. The PCR product was purified and BigDye terminating chemistry used to cycle sequence the DNA. The reactions were cleaned and sequenced on an ABI 3730XL DNA analyzer.
Correction of nucleotide sequences and assembly of contiguous sequences were performed using the computer program Sequencher 4.2.2 (Gene Codes Corporation. Ann Arbor MI). The data were partitioned into conserved and non-conserved regions based on the initial Sequencher alignment and then the alignment was refined using the program MAFFT (Katoh et al., 2005). The alignment was analyzed under parsimony.
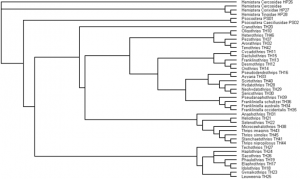
Parsimony (simplest assumption) analysis resulted in two most parsimonious trees, the strict consensus of which is presented. Nodal support values indicate that most nodes are well supported, though some of the deeper nodes within Terebrantia are lacking robust support. Thysanoptera is robustly resolved as a monophyletic with the suborders Tubulifera and Terebrantia each supported as monophyletic. Within the family Phlaeothripidae, the subfamily Idolothripinae is monophyletic while the Phlaeothripinae is paraphyletic. Within Terebrantia, the family Melanothripidae is placed as sister group to the remaining families. The family Aelothripidae is supported as monophyletic, but the family Thripidae is grossly paraphyletic across the Terebrantia. However, within Thripidae, some subfamilies are monophyletic including Panchaetothripinae.
This analysis includes a relatively modest sample of thrips diversity and only three genes, but the results are encouraging. A more thorough analysis including a broader selection of taxa combined with a wider range of genes is likely to provide additional insights into the phylogeny and evolution of this important insect order.
References
- Izzo, T. J., S. M. J. Pinent, and L. A. Mound. 2002. Aulacothrips dictyotus (Heterothripidae), the first ectoparasitic thrips (Thysanoptera). Florida Entomologist 85: 281- 283.
- K. Katoh, K. Kuma, H. Toh and T. Miyata (Nucleic Acids Res. 33: 511-518, 2005)
- Mound, L. A. 2005. Thysanoptera (Thrips) of the World–achecklist. http://www.ento.csiro.au/thysanoptera/worldthrips.html.
- Mound, L. A.and R. Marullo. 1996. The Thrips of Central and South America: An Introduction. Mem. Entomol. Int. 6:1-488.
- Mound, L.A. and D. C. Morris, D.C. 2003. The Morphological Background to Thysanoptera Phylogeny. Entomologische Abhandlungen Volume 61 No. 2: 151-152
- Shao, R., Barker, S.C. 2003. The Highly Rearranged Mitochondrial Genome of the Plague Thrips, Thrips imaginis. MBE Volume 20, No. 3: 362-370.
- Swofford, D. 2002. PAUP* Phylogenetic analysis using parsimony and other methods. Sinauer Associates. Sunderland, MA.
- Sorenson M.D. (1999) Treerot. Boston University, Boston.
- Wheeler, W.C., M. F. Whiting, Q. D. Wheeler, and J. Carpenter. (2001). Phylogeny of the extant hexapod orders. Cladistics 17: 1 89.
