Adam S. Frost and Professor David D. Busath, Zoology
Gramicidin A is a 15-amino acid peptide produced by several strains of the bacteria Bacillus brevis. The primary sequence of gramicidin A is HCO-L-Val-Gly-L-Ala-D-Leu-L-Ala-D-Val-LVal- D-Val-L-Trp-D-Leu-L-Trp-D-Leu-L-Trp-D-Leu-L-Trp-NHCH2CH2OH. The side-chains alternate chirality, with the odd-numbered residues having the usual L-chirality and the evennumbered residues having D-chirality. The alternating chirality of the side-chains forces them to project away from the lumen of the helix, yielding a tight wind with a central hole formed by the space within the helix. The secondary structure of gramicidin is a single-stranded, left-handed, b6.5-helix that dimerizes head-to-head (N-terminus to N-terminus) in lipid bilayer membranes to form a monovalent, cation-selective channel.1
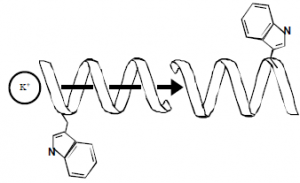
Gramicidin was the first identified and structurally characterized ion channel. While it is structurally simpler than channels in humans, gramcidin shares several features in common with these important channels. Like the potassium “shaker” channel, the carbonyl oxygens of the peptide backbone form the permeation pathway in gramicidin. Like the sodium channel, gramicidin is selective for monovalent cations, distinguishing between ions as similar in size as Na+ and Ca2+. Because its structural features are well known and easy to manipulate, the pore formed by the gramicidin molecule is ideal for studying ion channel structure-function correlations.
Eyring transition state theory and electrodiffusion continuum theories are both used to explain and predict the properties of biological ion channels. Debate continues over the strength and weaknesses of each of these theories. In this study, we evaluated the predictive power of a fourstate kinetic model to analyze K+ currents through the gramicidin A channel and eight analogs of gramicidin with fluorinated tryptophan side-chains. Nonlinear least-squares fitting of a 38 parameter space was used to assign rate constants for the current-voltage relations of gramicidin A, (5F-Indole)Trp9, 11, 13, and 15 gramicidin and (6F-Indole)Trp9, 11, 13, and 15 in two types of bilayers. The data set consisted of 1280 points, each one being the mean of three independent measurements of the single-channel current for each peptide in different concentrations of KCl and with different membrane potentials.
Eyring rate theory predicts that the rate constant for a given step in a reaction is exponentially related to the height of the energy barrier according to the relation:
![]()
In this equation, A is the rate constant, a is the activity of the permeating ion, DG is the height of the energy barrier, and a is the voltage dependence for the transition.2
Most attempts to model gramicidin current by means of Eyring rate theory divide the permeation process into three discrete steps. First, the ion diffuses up to the entry of the channel, loses its shell of hydrating waters, and binds to the carbonyl oxygen atoms of the channel backbone. Second, the ion jumps from its binding site at the entry of the channel to a symmetrical binding site at the exit of the channel (>20 Å away). Third, the ion escapes from the exit-side binding site and diffuses away from the channel into the bath. We refer to this model as the 3-barrier 2-site (3B2S) model because the potential energy profile contains three barriers, one for each of the steps described above, and two potential wells representing the symmetric binding sites. Five rate constants describe the allowed transitions between the four states: A, for ion entry into an empty channel; B, for ion exit from a singly occupied channel; K, for ion translocation from one binding site to the other; D, for ion entry into a singly occupied pore; and E, for ion exit from a doubly occupied channel.
The parameter set in the table shown below and the current-voltage (I-V) plots represent the best-fit. The I-V fit is visually good. However, The c2 statistic (reduced c2 = 8.56 with 1231 degrees of freedom) indicates that the probability is less than 1% that the Eyring rate theory predictions belong to the same population as the actual data.
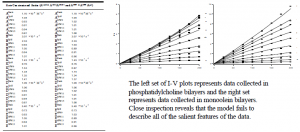
We concluded that the simple 3B2S kinetic model is too simple to accurately describe all of the data features observed for gramicidin conductance. Specifically, we have shown that the 3B2S model can not describe the diffusion-limited entry step (rate constant A), nor can it accurately predict the voltage dependence of the translocation step (rate constant K).
References
- Ketchem, R. R., Roux, B., and Cross, T. A. (1997) Structure. 5, 1655-1669.
- Eyring, H. (1936) J. Chem. Phys. 4, 283–291.
