Trevor Huff and Erin Bigler, Department of Psychology
Introduction
In this era of the human connectome, automated image analysis techniques, and large scale multi-site neuroimaging databases examining neuropsychological outcome across a broad spectrum of neurological and neuropsychiatric disorders, there is a particular need to address how to combine neuroimaging studies that use different volumetric sequences or Magnetic Resonance Imaging (MRI) studies performed on different platforms. Currently, there is a large volume of studies that utilize data from multiple scanning locations. This data, while important, cannot reliably be used for comparisons without taking into account the various differences that exist between MRI equipment. The purpose of this study is to investigate possible solutions to this existing problem and propose a direction for future investigations.
Methods and Results
In this study, two different approaches were examined. (1) Individual subjects were scanned on up to 4 different platforms at different intervals or (2) the same individual was scanned on the same scanner but with slight variation in the volumetric sequence. After these individuals were scanned, their images were then run through a tissue-based standardization protocol. In order to standardize the intensities of each sequence, we used a combination of fslmaths, Atropos segmentation (from the ANTs pipeline), and bet (Brain Extraction Tool). Once this was completed, FreeSurfer, a software packaged used to analyze MRI data, was used to make volumetric comparisons of different brain structures. FreeSurfer outputs were compared between both the standardized and non-standardized human phantom images. Then, statistical analyses and standardization were performed on a large database of MRI images (SOBIK). Both the nonstandardized and standardized groups from this dataset were analyzed and then compared.
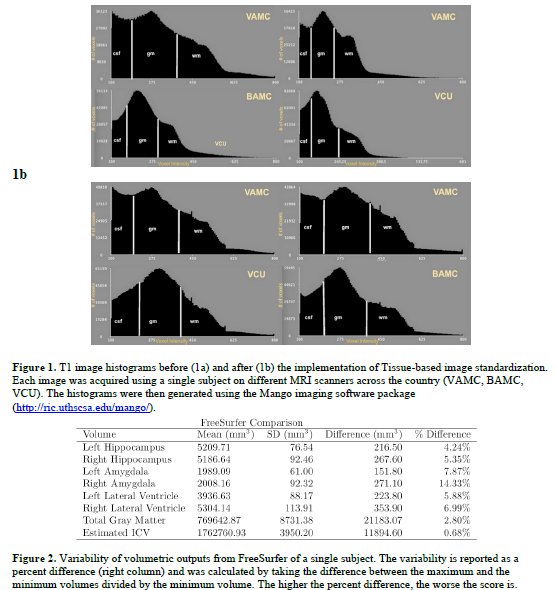
As shown in Figure 1, there is a noticeable difference in voxel intensities between the images acquired of a single subject at different locations across the country. In addition, the volumetric outputs provided by FreeSurfer also showed that there was great variability between the images of a single subject (see Figure 2). After tissue-based standardization, the FreeSurfer data remained greatly variable for the human phantom outputs. Statistical analyses show subtle differences between standardized and non-standardized images from the SOBIK dataset when morphological maps are run with different behavioral variables. In some instances, the standardized images show a stronger result or remove presumably false results from the OI population.
Summary and Conclusion
The human phantom data suggests that standardization does little to correct the underlying problems with multi-site imaging. Improvements can be made to more closely match individual histograms, but this does not improve the replicability of volumetric MRI outputs for multi-site studies. In addition, there is no evidence to suggest that image standardization is beneficial in any way for these kinds of investigations. More importantly, there is an increased need to find the source of variability and propose alternate ways to reliably compare large datasets such as the ones provided in this study.