Julianne H. Grose
Evaluation of academic objectives
The academic objectives of this MEG were to train undergraduate and graduate students on the scientific process while providing valuable insight into the genetic factors that may play a role in the development of diabetes in the Wasatch front population. We were able to meet these objectives as evidenced by the training of two graduate students and seven undergraduates and by the identification of a novel genetic allele which most likely contributes to diabetes.
Articles in preparation resulting from this MEG
Demille et al. Identification of a Novel HNF1beta Allele Associated with Familial Early Diabetes in a Utah Family. In preparation
Evaluation of the mentoring environment
We had weekly lab meetings where the students took turns presenting their findings, providing students with a broader view of science and aiding in the development of presentation skills. In addition, weekly project-specific subgroup meetings were formed where data interpretation, experimental theory and protocols, and solutions to any current experimental difficulties were discussed. Students were trained by hierarchies within these subgroups, those who have the most experience were involved in training the newer lab members. This hierarchy facilitated higher learning as students explained concepts behind their research to others and gradually acquired more independence and responsibility. All students were encouraged to take ownership of their project and to contribute by offering new avenues to explore as well as alternative experimental techniques.
Undergraduates trained in the Grose lab in 2013-2014 sponsored by this MEG:
- Evaneline Taylor (patient recruitment and patient advocate)
- Katie Harris (patient recruitment)
- Benjmin Donovan (literature searches and help with DNA sample prep)
- Sined P. Yepez (literature searches and help with DNA sample prep)
- Wesely Goar (literature searches) Jared Adnerson (certified phlebotomist and literature searches)
- Emily Bennion (certified phlebotomist and literature searches)
- Graduate students trained in the Grose lab 2013-2014 sponsored by this MEG: Desi DeMille (DNA sample preparation)
- Kai Li Ong (Mutation verification)
Description of the results/findings of the project
Aim 1: Identify families in our local community who appear to have genetic factors contributing to the development of diabetes.
Local doctors report a high number of diabetic patients with several immediate family members affected. Our goal was to identify patients with an apparent familial genetic factor for further study. I consulted wth Dr. Keoni Kauwe, an expert on identifying complex and rare genetic factors contributing to disease, and Dr. Laura Bridgewater and together we established the following criteria for patients to participate in this study. First, a patient must have been diagnosed with diabetes before the age of 40. Second, they must have at least two other relatives that have been diagnosed with diabetes (parent, sibling, child, aunt, uncle, first cousin, niece or nephew). Finally, they must be willing to provide a family pedigree and donate a blood sample for DNA analysis. Flyers announcing the study were placed in clinics along the Wasatch front and patients have contacted our study via email. Interested patients then came to BYU where the details of the study are explained by a research scientist (graduate student Desi DeMille or myself) and the family pedigree is obtained. Blood samples were then acquired by a certified phlebotomist (undergraduates Emily Bennion or Jared Anderson). Pedigrees and blood samples were randomly numbered with pedigrees stored in an office under lock and key and blood samples stored in the lab. We obtained blood samples and pedigree charts from 12 individuals.
Aim 2: Isolate and sequence whole genomic DNA from five unrelated individuals with the greatest apparent familial predispositions.
Due to the cost of human whole genome exome DNA sequencing, we were limited to sequencing five samples from the individuals whose family pedigree appeared to be the most promising for identifying a novel contributing factor. All five whole genome exomes were sequenced. From the five samples, one had a novel mutation in HNF1α. Two had known mutations in genes that have previously been shown to associate with diabetes, and the other two had no obvious genetic factors.
Mutations in the HNF1α or HNF1β gene are associated with several types of diabetes mellitus, with dominant mutations resulting in MODY3. Common variants of HNF1α or HNF1β are associated with type 2 diabetes, including I27L in exon 1 (Holmkvist et al., 2008; Holmkvist et al., 2006; Urhammer et al., 1997). HNF1 α and β are transcription factors that influence the development and function of the pancreas, kidney and liver, along with several other organs. They specifically target several genes in involved in β-cell function including, insulin, glucose transporter 2, hepatocyte growth factor activator pyruvate kinase, collectrin and HNF4α (Evans et al., 2008; Ladias et al., 1992; Odom et al., 2004). Thus, disruptions in HNF1 impair glucose-stimulated insulin secretion by pancreatic β cells, resulting in diabetes. In addition, certain HNF1 β mutations have been linked to multiple types of renal disease, kidney atrophy. kidney malformation, genitourinary tract abnormalities, and atrophy of the pancreas. The MODY phenotype is observed in the majority of patients with HNF1B mutations (approximately 50-70% of patients). MODY is often misdiagnosed as either type I or type II diabetes due to the infrequency of genetic testing for diabetic patients. There are many different causes for MODY which lead to varying forms and degrees of the disease requiring more specialized treatment. MODY 3 patients are often misdiagnosed as type 1 Diabetics while they are sensitive to low-dose sulfonylruea and do not usually need insulin treatment (Pearson et al., 2003; Shepherd et al., 2003), making proper diagnosis key to proper treatment. Thus, the identification of a novel HNF1β mutation in a Utah patient with familial diabetes is most likely the causative factor.
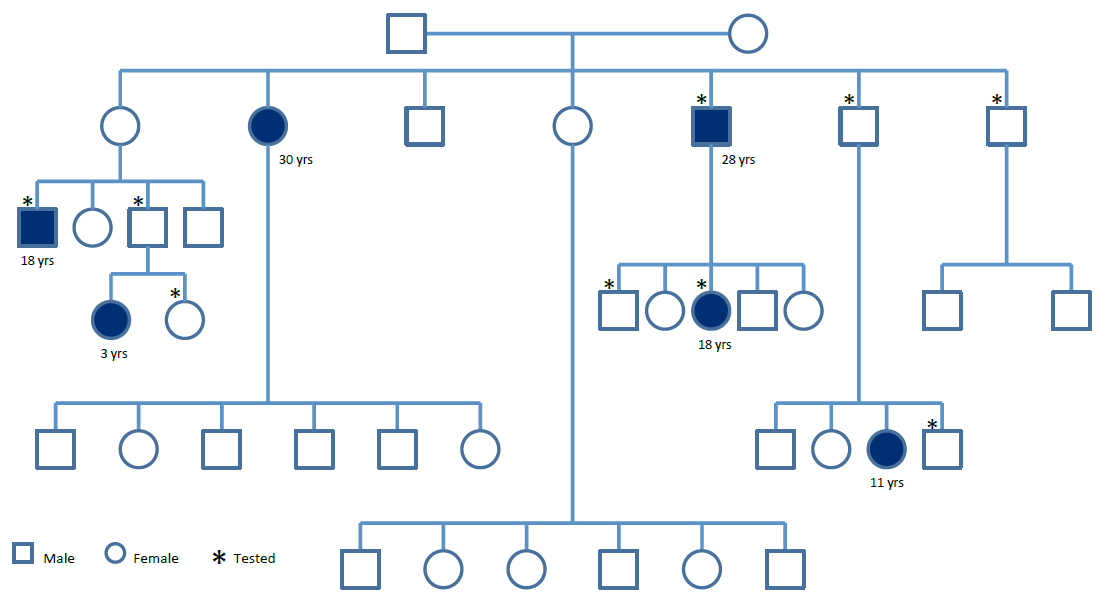
Aim 3: Identify novel genes contributing to diabetes by performing whole genome exome sequencing on two families with the greatest familial predispositions. Instead of focusing on two families, we decided to devote our time and effort towards this newly identified HNF1B allele. We, therefore, collected DNA samples from many other family members in order to verify the absence or presence of this mutation. The familial pattern for diabetes and the samples we collected are indicated in Figure 1. We have all of the samples and are still verifying the sequence of the family members, since we obtained unexpected the first time.
Figure 1. Pedigree of a family with familial diabetes. The years is the age diagnosed with diabetes.
Conclusions: We began this work due to the passion and influence of a BYU undergraduate, Evangeline Taylor. Unaccustomed to this type of research, I underestimated the man-power and funding it would take. Despite this setback, I am hopeful we will be able to confirm the HNF1β mutation and write a scientific article on our findings in the next few months. We are grateful for the assistance of Dr. Keoni Kauwe in the genome analysis.
Description of how the 2013-2014 budget was spent
- $9,600 5 undergraduate for 15 weeks, 10 hours per week at $8.25 per hour.This pay was for months (spring/summer) in which these students were not be enrolled in the research for credit course 494R.
- $3,000 pay for 1 masters student (Kai Li Ong). 30 weeks, 10 hours per week, $10 per hour.
- $2,400 Research expenses include laboratory consumables, reagents and supplies for wet lab research.
- $700- DNA extraction/prep kit for DNA sequencing
- $600- phlebotomy supplies (tubes/gloves/sterile pads, etc)
- $1000- basic lab supplies gloves, pipettes, microcentrifuge tubes, kimwipes, parafilm, supplies for PCR and other consumables
- $5,000 DNA sequencing costs–Whole genome exome sequencing for 5 individuals (~$850 each) plus BYU DNA sequencing center costs for PCR amplification and sequencing.