Background and Significance
Neofunctionalization is the process by which genes gain a new or modified function as a result of gene duplication and modification of one or more of the new genes. This exciting area of research helps explain the diversification of protein functions in cells. The number of documented examples has increased in recent years in all classes of organisms (Deng et al., 2010; Han et al. 2011; Cannela et al. 2014; Sakuma et al. 2013; Zhang et al. 2014). In some cases a protein that was previously thought to function in only one cellular compartment has been shown to also function in another compartment with a modified or new function. This includes proteins localized to mitochondria and chloroplasts.
The divergence between organelle genomes of animals and plants during evolution is profound and provides different scenarios for neofunctionalization. The concept that organellar DNA maintenance proteins serve as a rich source for protein neofunctionalization studies was first suggested by observations by the Butow group in surveys of mitochondrial nucleoid proteins in yeast (Kaufman et al. 2000). They subsequently identified the mitochondrial aconitase Krebscycle enzyme as being essential for mitochondrial DNA (mtDNA) maintenance independent of its catalytic activity, and this protein may integrate metabolic signals with mtDNA maintenance (Chen et al. 2005). A whole host of proteins of already known function were now found to have DNA binding properties. Another group showed that yeast aconitase plays important functions in different metabolic pathways in the cytosol and mitochondria (Regev-Rudzki et al. 2005). Human mitochondrial transcription factor A, which is essential for mitochondrial gene expression, also functions in the cell nucleus to regulate cancer cell growth (Han et al. 2011).
Plant organellar MSH1 has been shown to play a role in mitochondrial and chloroplast genome maintenance and to influence plant growth and stress response via plastid-induced epigenetic modulation (Shedge et al. 2007; 2010; Xu et al. 2011; 2012). Neofunctionalization of a number of other plant organelle dual-targeted proteins has been proposed and shown to affect organelle gene expression (Babiychuk et al. 2011; Brandao and Silva-Filho 2010) or organelle membrane protein function (Topel et al. 2012). In each of these cases a gene duplication event has occurred and one of the gene copies underwent sequence changes that have affected regulation of gene expression or contributed to a new function for the protein. The evolution of dual-targeted proteins has been proposed to contribute to neofunctionalization (Xu et al. 2013).
Chloroplast DNA (ctDNA) and mtDNA are replicated by nuclear-encoded proteins that are imported into the organelles. The major proteins involved in plant organelle DNA replication including two DNA polymerases, DNA primase/helicase, single-stranded DNA binding protein (SSB), and some accessory proteins have been identified. Many are present in two copies and have been shown to be dual-targeted to chloroplasts and mitochondria (Cupp and Nielsen 2014; Moriyama and Sato 2014; Gualberto et al. 2013). DNA and amino acid sequence analysis has identified stretches of additional amino acids within the coding regions of Arabidopsis SSB that may provide new or altered functions or interactions with other proteins. In addition, we have identified different upstream promoter elements for the DNA PolIA and PolIB genes, and their expression may not be regulated in the same way. These additional sequences are highly conserved in plants, suggesting that they may be important for function. However, the expression and biochemical properties for most of the replication proteins have not been studied. A number of factors may be important in the regulation of expression of proteins involved in organelle DNA replication. Light, stress and other environmental conditions appear to contribute to regulation of organelle genome replication and copy number. Light-regulated organelle DNA replication is complex and is regulated differently in land plants compared to algae (Moriyama and Sato 2014). Reactive oxygen species are involved in regulating mtDNA replication in yeast (Hori et al. 2009), and ctDNA replication in algae is regulated by cellular redox state and light (Kabeya and Miyagishima 2013; Ohbayashi et al. 2013).
Based on sequence homology many of the dual-targeted organelle DNA replication proteins appear to have the potential to provide redundancy; i.e. if one gene is mutated the other could compensate to provide the needed function and allow continued growth. However, analysis of mutants in several of these genes shows that there is some disruption of plant development and growth when one of the genes is mutated, strongly suggesting that they are not fully redundant (see Preliminary Studies). Because plant organelle DNA replication machinery has been poorly studied this provides a rich area for identifying new examples of neofunctionalization of proteins that have occurred after endosymbiosis.
Plant organelle DNA polymerases. All sequenced plant genomes encode two highly conserved organelle-targeted DNA polymerases (Moriyama and Sato 2014; Moriyama et al. 2011; Ono et al. 2007). These two enzymes have been termed DNA polymerase IA (PolIA) and IB (PolIB), and are the only organellar DNA polymerases. Initially PolIA was shown to localize exclusively to chloroplasts while PolIB is localized to both organelles (Elo et al. 2003). However, both were subsequently shown to be dual-localized (Carrie et al. 2009). Dual-targeting is observed for stable constructs that contain the native PolIA promoter, but the localization is affected by the upstream promoter region and depends on a potential alternate start codon (Christensen et al. 2005). It is possible that the dual-targeting of PolIA is developmental stage specific, but no detailed studies have been reported that analyze when each is expressed and where each is localized during plant development or in response to different environmental signals.
DNA polymerases are responsible for genome replication, but Parent et al. (2011) reported divergent roles of the organellar DNA polymerases in Arabidopsis, showing that DNA PolIB, but not PolIA, functions in ctDNA repair, as mutations in PolIB increase susceptibility to DNA damaging agents. Mutants in either gene alone retain viability but exhibit varying levels of reduction in growth and development (Cupp and Nielsen 2013; Parent et al. 2011), supporting the possibility of different functions for each. No gene co-expression data are available for PolIB in the ATTED database (http://atted.jp), but data are available for PolIA. PolIA is co-expressed with chloroplast localized RecA, helicases, and proteins involved in plant development. Mutants in DNA PolIA showed no susceptibility to DNA damage, and its role in organelle DNA replication and maintenance is unclear.
We have analyzed DNA PolIB mutants and observed a 30% reduction in mtDNA copy number and changes in respiration activity (Cupp and Nielsen 2013). The PolIB homozygous mutant has a large decrease in mitochondria size and a simultaneous significant increase in the number of mitochondria per cell. However, no effects on chloroplast size or number were observed. We are currently analyzing the DNA PolIA mutants to determine the effect of the mutation on genome copy number and organelle structure, and on respiration and photosynthesis activity. This protein is dual-localized under specific conditions (Christensen et al. 2005), but under other conditions PolIA is preferentially localized to chloroplasts (Elo et al. 2003, see Preliminary Studies), and it may play a more predominant role in chloroplasts and have different functions in each organelle.
Single-stranded DNA binding proteins (SSB). Another organelle replication protein with potential for evolved and/or multiple functions is SSB1 (single-stranded DNA binding protein). The Arabidopsis SSB1 protein is highly conserved with SSB from bacteria and other plant species, but it has an N-terminal extension of 68 amino acids not present in the bacterial protein (Edmondson et al. 2005). The first 23 amino acids is predicted to be an organelle localization signal that gets cleaved off upon import into mitochondria, leaving a 45 amino acid section with unknown function. These 45 amino acids are highly conserved across plant species and are also found in mosses but not in the bacterial or human mitochondrial orthologs or in other organisms (see Preliminary Studies). However, in another Arabidopsis SSB homologue, SSB2, there is a similar extra stretch of amino acids in the same region compared with bacterial SSB but with a different sequence than SSB1. SSB1 also has a unique PDF motif in the C-terminal end of the protein (Zaegel et al. 2006). Analyses of these extra regions by pFAM, InterPro and other protein domain prediction programs do not suggest any potential function, but the soybean orthologue of Arabidopsis SSB1 has previously been shown to bind to the ctDNA replication origin sequences (Lassen et al. 2011). The additional amino acids may be involved in interactions with other replication proteins. No co-expression data is available for SSB1 in the ATTED database.
Chloroplast genomes and ctDNA replication. Chloroplast genomes (ctDNA) range in size between 130–160 kbp in higher plants (Palmer 1985). CtDNA is associated with replication and compacting proteins in nucleoids (Majeran et al. 2012). Displacement loop (D-loop) replication origins have been mapped in ctDNA near the rRNA genes in several species (Kunnimalaiyaan and Nielsen 1997). Rolling circle replication may occur after completion of D-loop replication (Nielsen et al. 2010). CtDNA may replicate by more than one mechanism specific to the developmental stage, similar to some bacteriophages (Nielsen et al. 2010; Muhlbauer et al. 2002). The early rapid expansion of ctDNA may be facilitated by rolling circle or recombinationdependent DNA replication (RDR, Nielsen et al. 2010; Scharff and Koop 2006; Oldenburg and Bendich 2001; 2003), followed by a maintenance level of replication in older tissues (Nielsen et al. 2010). Some research has suggested that as cells and tissues age ctDNA levels decrease and become more fragmented (Kumar et al. 2014).
Mitochondrial genomes and mtDNA replication. Mitochondrial genomes vary widely in size, ranging from a very compact 16,500 bp in humans and other vertebrates to 30,000-90,000 bp in yeast and other fungi, and from 208,000 to about 2,000,000 bp in higher plants (Knoop 2004). The much larger size and different genome structure and organization in plants indicates a profound divergence of mitochondrial genome evolution between plants and animals. The major structure of plant mtDNA exists as subgenomic molecules that are primarily linear (reviewed in Bendich 1993 and Backert et al. 1997; Oldenburg and Bendich, 2001), suggesting that multiple molecules must be coordinately replicated to ensure maintenance of essential genes, which are dispersed throughout the genome. During maize development mtDNA becomes fragmented and copy number decreases (Kumar et al. 2014). Because of the size and complexity of the genome, the mechanism for plant mtDNA replication is still not clear. In plants and fungi it has been suggested that mtDNA may replicate by RDR and/or a rolling circle mechanism similar to bacteriophage T4 DNA replication (Backert and Borner 2000; Oldenburg and Bendich 2003; Cupp and Nielsen 2014), which is much more complex than replication of animal mtDNA (Bogenhagen and Clayton 2003). It has been determined that on average some mitochondria contain less than a full genome equivalent (Kanazawa et al. 1994), likely due to frequent fission and fusion of plant mitochondria (Arimura et al. 2004). These observations raise questions concerning how plant mtDNA replication and distribution occurs to ensure that each new mitochondrion receives a full functional copy of the genome.
Recombination-dependent replication (RDR). With the growing evidence that RDR may be a major mechanism for DNA replication in one or both plant organelles, this infers that one or more recombinases may be involved to facilitate DNA synthesis. Three bacterial RecA orthologs have been identified as RecA1, RecA2, and RecA3 in the Arabidopsis genome (Khazi et al. 2003; Shedge et al. 2007). RecA1 is localized to chloroplasts, RecA2 is dual-localized, and RecA3 is found in mitochondria. As mentioned above, RecA1 is co-expressed in Arabidopsis along with DNA PolIA, supporting the possibility that these two proteins function together. RecA1 and RecA3 are coexpressed with different genes involved in development (ATTED database). No coexpression data is available for RecA2. Shedge et al. (2007) reported that RecA1 is essential in Arabidopsis, as homozygous mutants are not viable. If this protein only functioned in repair it would be expected that mutants would still be viable but unable to repair DNA damage, which suggests that RecA1 may play a critical role in ctDNA replication, and not just in DNA repair. In support of this, the involvement of RecA1 in maintenance of ctDNA integrity has been reported (Rowan et al. 2010). Mutants in RecA1 resulted in altered ctDNA structure and reduced ctDNA levels, along with an increase in single-stranded regions of ctDNA. Parallel functions for RecA2 and RecA3 may contribute to mtDNA replication and genome maintenance. Strand invasion catalyzed by one of the RecA homologs followed by extension of new DNA synthesis by one or both of the organellar DNA polymerases may be directly involved in organelle genome replication.
Previous work from our laboratory. Research in my laboratory has focused on plant ctDNA and mtDNA replication and recombination, including proteins and mechanisms involved. We have published numerous papers in this area, including characterization of several ctDNA and mtDNA replication proteins (RecA, SSB1, DNA PolIB, a potential ctDNA origin-binding protein (Lassen et al. 2011), Twinkle helicase/primase, ctDNA primase, and topoisomerase I) and characterization of ctDNA replication and mtDNA recombination (details in reviews by Cupp and Nielsen 2014; Nielsen et al. 2010; Kunnimalaiyaan and Nielsen, 1997). Surprisingly, despite an abundance of available information on plant organelle genomes and gene expression (Schuster and Brennicke 1994; Kunnimalaiyaan et al. 1997; Mackenzie and McIntosh 1999), the mechanisms that control organelle DNA replication and genome copy numbers are not well understood. Organelle DNA levels vary significantly (Kumar et al. 2014) in different ages and types of tissues depending on energy needs and location in the plant, indicating that regulation of mtDNA and ctDNA replication and genome copy number occurs. This regulation is distinct from known mechanisms that control replication of bacterial and eukaryotic nuclear genomes, which suggests either the involvement of additional unique proteins in plant organelles or that neofunctionalization of existing proteins has occurred.
Central hypothesis and objectives of this proposal. The central hypothesis of this study is that organelle-localized SSB1 and the two DNA polymerases have undergone neofunctionalization so that each plays a distinct role in ctDNA and/or mtDNA replication and genome maintenance. Each of these genes contains sequences that are highly conserved in plant orthologues but are unique to plants. By studying the localization, spatial and temporal expression and protein interactions of these proteins we plan to characterize their functions in organelle genome maintenance and the nature of evolutionary changes they have undergone. Specific objectives of this project include:
- Analyze the Arabidopsis organellar DNA polymerase and SSB1 genes for sequences that may be responsible for new or altered functions and examine their interactions with other components of the DNA replication machinery by co-immunoprecipitation.
- Conduct detailed expression analysis of each replication protein in different plant tissues and stages of growth in wild-type and mutant plants using GUS assays and confocal microscopy with GFP- and RFP- gene fusion constructs to understand which proteins are coexpressed under different conditions.
- Develop model(s) for evolution of neofunctionalization of organelle DNA replication proteins and mechanism(s) for plant organelle DNA maintenance.
Preliminary Studies
During a recent professional development leave in the laboratory of Dr. Sally Mackenzie at the University of Nebraska (June-Oct. 2014) I was able to construct native promoter and 35S promoter-GFP, –RFP and -GUS fusions for the DNA polymerase, SSB1, and Twinkle proteins. The native promoter gene fusion constructs contain 1.5 kbp of upstream promoter region and the entire coding region fused to the reporter gene coding region, which allow determination of localization of each protein by confocal microscopy. Because of the native promoter, expression levels may be low and the proteins may not be expressed in all tissues or all stages of development, so constructs that use the CaMV 35S promoter from pCAMBIA 1302 to drive constitutive expression have also been made. Each of these constructs include the coding region for the entire replication protein fused in-frame with the reporter gene. Some of these constructs have already been analyzed in transiently transformed plants (see Figs. 1 and 5).
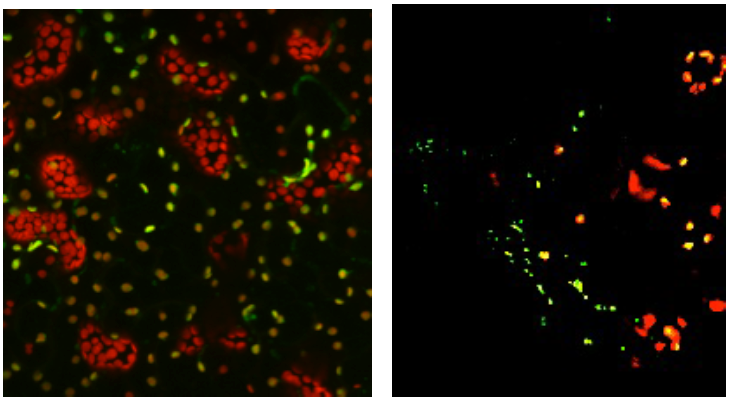
Organellar DNA polymerases. We have characterized mutants in DNA PolIB and shown that PolIB plays a significant role in mtDNA replication (Cupp and Nielsen 2013). Expression data indicates that PolIA is coexpressed with chloroplast-localized RecA, regulatory genes, and proteins involved in meristem development, root growth, and seed dormancy (http://atted.jp). This suggests involvement of PolIA in ctDNA replication and in plant development. No coexpression data for PolIB is available in the ATTED database. Transient expression analysis of fusion proteins has indicated that localization of DNA PolIA appears to be more pronounced in chloroplasts (similar to what was observed by Elo et al. (2003), while PolIB is more pronounced in mitochondria (Fig. 1). These images were generated at the Univ. of Nebraska plant imaging facility during my leave. We acknowledge that transient expression assays may not accurately reflect the true protein localization, so stable transformants using the floral dip method (Clough and Bent 1998) expressing the fusion proteins have been generated and are now being screened.
We have recently initiated analysis of PolIA mutants, which exhibit a slight reduction in development compared to wild type, less than with the PolIB mutants (Cupp and Nielsen 2013). We have observed a reduction in organelle genome copy number in the mutants for either gene compared to wild type (Cupp and Nielsen 2013 and unpublished data for PolIA). The ability of mutants in either gene to reach maturity and produce seeds suggests partial redundancy, so that when one gene loses function the other gene can only partially compensate. We have been unable to obtain mutants homozygous for insertions in both genes, as expected since at least one copy of the DNA polymerase is essential for genome replication. Therefore, we examined the coding and promoter regions for both genes. While the two DNA polymerases are very similar at the coding sequence level, the genes have very different upstream promoter regions, indicating that expression may be regulated by distinct elements (Fig. 2). In addition, there are differences in the nuclease domain of the protein-coding region although the two appear to have arisen by gene duplication (Fig. 3). PolIB but not PolIA has been shown to be involved in ctDNA repair (Parent et al. 2011), although localization of this protein is more pronounced in mitochondria (Fig. 1) and mutants affected mtDNA but not ctDNA copy number (Cupp and Nielsen 2013). These observations support the possibility that organelle-localized DNA polymerases may have undergone neofunctionalization to provide new functions specific for each in organelle DNA replication and maintenance.
Figure 1. Localization of the DNA polymerases in transiently transformed plants. DNA PolIA is primarily localized to chloroplasts (left) while DNA PolIB is localized to mitochondria (right). The constructs have the full-length DNA polymerase gene fused with RFP (PolIA) or GFP (PolIB) to allow visualization by confocal microscopy. Chloroplasts autofluoresce red under these conditions; RFP and GFP emit fluorescence at a different wavelength and their presence is indicated by the green color. Mitochondria are much smaller than chloroplasts (small green spots in the panel at right). Note the punctate pattern of green in many of the chloroplasts, suggesting that the enzyme is associated with DNA in the nucleoids.
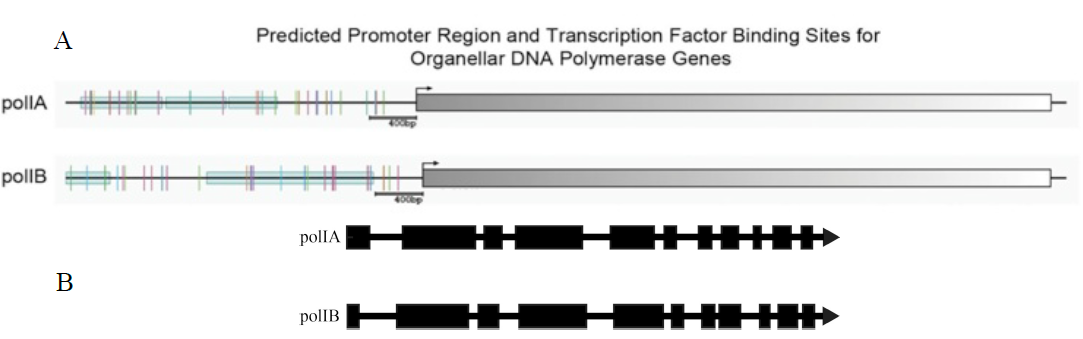
Figure 2. A. Predicted promoter elements of the DNA polymerase genes. The colored lines upstream of the coding regions indicate distinct elements, showing that the two genes have dissimilar upstream control regions and thus have the potential to be differentially expressed. B. Exon-intron structure of the two genes is highly conserved, characteristic of duplicated genes.
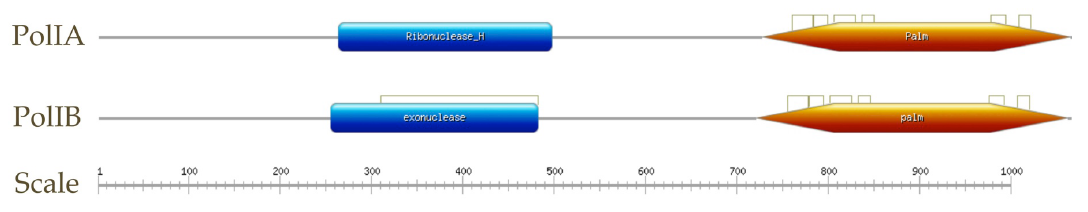
Figure 3. Functional domains of PolIA and PolIB. The two proteins have 71.3% protein identity in the polymerase palm domains (catalytic site colored orange), but have vary in the N-terminal region of the proteins, including the nuclease domain (marked in blue).
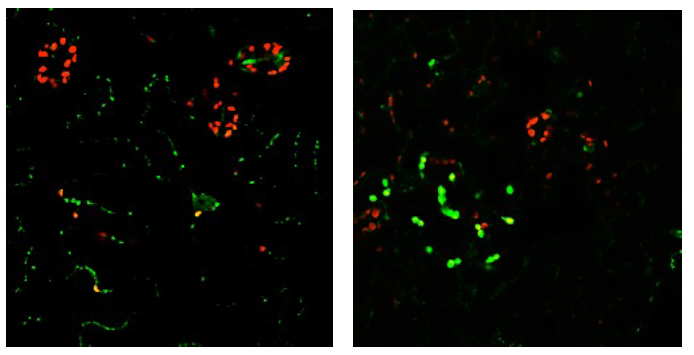
Single-stranded DNA binding protein (SSB). A T-DNA insertion in the 5’UTR region (SAIL_378_E03) of SSB1 has an unusual phenotype, with very slow growth when plants are grown in 16 hr light/8 hr dark conditions. Germinating plants are light yellow and young seedlings remain pale compared to wild-type plants (Fig. 4 left panel). The mutants eventually turn green and produce seeds. However, when the mutant is grown under 12 hr light/12 hr dark conditions the plants grow at nearly the same rate as wild type (Fig. 4 middle panel). All plants grow more slowly in 12 hr light, but in these conditions the mutant appears to be able to adjust and grow more like wild-type. The mutant remains somewhat pale until it begins to put up shoots, when the plants become as green as wild-type (Fig. 4 right panel). The pale phenotype of the SSB1 mutant suggests a chloroplast effect, but the protein was initially reported to localize only to mitochondria (Edmondson et al. 2005). However, our analysis of a native promoter-full gene-RFP construct indicates chloroplast localization in some cells (Fig. 5), supporting a potential role in both organelles. It may be that chloroplast localization occurs only under some conditions. Interestingly, we have found that the SSB1 ortholog in soybean binds specifically to the ctDNA D-loop origin sequences (Lassen et al. 2011). It is possible that SSB1 plays a role in initiation or regulation of ctDNA replication, and may interact with other replication proteins in both organelles. As part of this study we will characterize interacting proteins identified by coimmunoprecipitation and yeast two-hybrid screening (See Experimental Plan).
Figure 4. Growth of SSB1 mutants in 16 hr light/8 hr dark (panel at left) and 12 hr light/12 hr dark (panels in the middle and right) conditions. The orange tabs in the left panel mark the SSB1 mutants, blue indicates a different mutant, and white indicates wild type. In the middle panel the yellow marker indicates the SSB1 mutant plants, which still have a yellowish center compared to normal plants at top (red marker) though both sets are about the same size. However, as the same plants get older and prepare to put up shoots (at right) they become green.
Figure 5. SSB1-RFP (native promoter construct for both panels) is predominantly localized to mitochondria (punctate green points in the image at left) but has been observed to localize to chloroplasts in some cells (green chloroplasts in the right panel).
Alignment of the SSB1 full-length amino acid sequence with orthologs from other plants and with SSB sequences from bacteria and eukaryotic mitochondria shows a significant stretch of Nterminal amino acids that is present in Arabidopsis SSB1 but is absent in Arabidopsis SSB2 and SSB proteins from other organisms (Edmondson et al. 2005; Fig. 6). While only a few sequences are included in this alignment, it is clear that the plant homologues of Arabidopsis SSB1 show a high degree of sequence conservation, suggesting that this region may play an important role in the function of the protein. It may play a specific role in interactions with other proteins and/or regulation. In support of this, we found that the soybean ortholog of this protein binds sequencespecifically to the double-stranded ctDNA ori region rather than only binding single-stranded DNA (Lassen et al. 2011). The potential for multiple roles for SSB1 and interactions with other replication proteins will be studied as part of this project (see Experimental Plan).
Figure 6. Alignment of SSB protein sequences. The two plant organelle-localized SSB orthologs have an extended N-terminal region but with different sequence, while lacking some amino acids at the C-terminal end compared to bacterial and human mitochondrial SSB.
Experimental Plan
We have published or recently been trained with all of the methods outlined in this section. I completed a sabbatical leave in Sally Mackenzie’s lab at the Univ. of Nebraska in 2014 where I was able to construct several of the gene fusion constructs that will be used. My students at BYU are currently analyzing these constructs.
Sequence and phylogenetic analysis. During my sabbatical leave in Sally Mackenzie’s lab I created reporter gene constructs for analysis of expression and localization of several organellar DNA replication proteins. I also initiated sequence analysis to identify unique sequences that will be further analyzed as follows. The sequence of each protein will be further analyzed by pFAM (http://pfam.xfam.org/), Uniprot (http://www.uniprot.org/), NCBI/BLAST (http://blast.ncbi.nlm.nih.gov/) and other resources to identify unique regions and predict potential functions or interactions with other proteins. Phylogenetic analysis of each protein will be conducted to determine the conservation and relationship with proteins from other species (Moriyama et al. 2014; Moriyama and Sato 2014). Sequences of replication protein orthologs will be obtained from Genbank and used to generate alignments and phylogenetic trees will be constructed by the Maximum Likelihood method using the TreeFinder software (Jobb et al. 2004). The analysis will also be done with specific regions of the proteins, such as the Nterminal sequence of SSB1 that is unique to plants (Fig. 6), similar to what has been done for functional domains of DNA polymerase I and the 5’-3’ exonuclease domain of plant organellar DNA polymerases (Moriyama and Sato 2014). This will provide insights into the degree of conservation of the plant-specific regions of the proteins, which will be tested for interactions with other molecules.
Co-immunoprecipitation and protein identification. The fusion constructs that we have developed will be used for co-immunoprecipitation (co-IP) analysis (Fabregas et al. 2013) to identify proteins that bind to the DNA polymerases and SSB1 using antibody against GFP that is present in the gene fusion constructs we have made. Recovered proteins will be prepared for mass spectrometry using the FASP protocol (http://openwetware.org/wiki/Prince:FASP), boiled to reduce phosphatases and kinases, and sonicated to break DNA (Wiśniewski et al. 2009). Trypsin digested peptides will be analyzed using the Orbitrap mass spectrometer in the BYU Chemistry & Biochemistry Dept. Peptides will be separated by strong cation exchange and reverse phase chromatography and analyzed by high mass accuracy/resolution mass spectrometry. Spectral matches to proteins in the UniProt Arabidopsis proteome will be made with SequestHT, Mascot, and MS-GF+ (Eng et al. 1994; Kim and Pevzner 2012). Percolator (Käll et al. 2007) will be used to reconstruct protein sequences and optimize peptide and protein statistical thresholds to maximize hits at a 1% peptide and protein false discovery rate. Data for any new proteins identified will be submitted to the appropriate database (AT_Chloro for chloroplast proteins at http://www.grenoble.prabi.fr/at_chloro/ and at www.plprot.ethz.ch, and the Arabidopsis mitochondrial protein database at www.plantenergy.uwa.edu/au/applications/ampdg/index.html).
As an additional or alternative approach we will utilize yeast-two hybrid screening to identify proteins that interact with each of the replication proteins we are working with. In addition, the regions of the proteins responsible for the interactions detected by co-IP will be further investigated by generating expression constructs that include specific regions of the initial protein bait for analysis by yeast two-hybrid screening. For example, if SSB1 is found to interact with other proteins, specific regions of the SSB1 coding region such as the first 68 amino acids prior to the DNA binding domain will be tested to determine if the interaction is specific to this region. Other overlapping segments of this gene coding region and the genes for the two DNA polymerases will also be examined to identify the precise interacting regions. Once these regions are identified we will determine if these sequences are conserved in other plants and organisms, and phylogenetic analysis of these specific domains will be done as described above. From the co-IP and yeast two hybrid analyses we expect to identify interactions with other already known replication proteins but we also may detect interactions with other proteins that may be involved in organelle genome replication or its regulation. This may include one of the RecA homologues if required for recombination-dependent replication. Any novel proteins that are identified will be analyzed for gene expression and localization relative to SSB1 and the DNA polymerases.
In addition, we will isolate nucleoids from chloroplasts and mitochondria and analyze the nucleoid proteins by mass spectrometry using the Orbitrap mass spectrometer to measure levels of each component of the organelle DNA replication machinery. Various plant tissues will be collected at weekly time points and organelles and nucleoids isolated by differential centrifugation and Percoll gradients (Weigel and Glazebrook 2002; Majeran et al. 2012; Huang et al. 2014). Protein spectral counts will be used to identify any proteins that show a significant increase or decrease in levels in the organelle nucleoids from the different samples. This will also be carried out for analysis of nucleoid proteins in mutant plants that are described below compared to wild-type. The proteomics analysis will not only help to identify the levels of specific DNA replication proteins in nucleoids and total organelle fractions in different tissues during different stages in plant development, but also in the SSB1 and DNA polymerase mutants versus wild-type plants. While the chloroplast and chloroplast proteome has been widely studied (Olinares et al. 2010; Taylor et al. 2009; 2011), the plant mitochondrial proteome is still not well understood (Pfalz and Pfannschmidt 2013). Lee et al. (2013) reported that less than 30% of the predicted mitochondrial proteins in Arabidopsis have been experimentally verified. Our work will likely confirm other mitochondrial proteins. While some proteins may be unstable or present at low levels, by isolating concentrated fractions of the organelles from specific tissues and from young plants we should identify DNA replication proteins and other organelle nucleoid proteins.
Confocal microscopy of reporter gene constructs. During my stay in Sally Mackenzie’s lab I successfully generated native promoter-reporter gene fusion constructs for the two Arabidopsis DNA polymerases, SSB1, Twinkle DNA primase/helicase, RecA2, and a few others, following previously published methods from the Mackenzie laboratory (Elo et al. 2003; Christensen et al. 2005). The GFP- and RFP-gene fusion constructs generated for the transient expression analysis (Figs. 1 and 5) have been used to create stable transformants by floral dipping (Clough and Bent 1998), and are now being screened. Confocal microscopy of stably transformed plants will be used to confirm the observations with transient transformants. Localization of each fusion construct (primarily the DNA polymerases and SSB1 for which was already have all of the needed constructs, along with other organelle replication proteins as time permits) in the stable transformants will be determined using the Olympus FluoView FV 300 confocal laser scanning microscope in the PDBIO Department at BYU or in collaboration with the Mackenzie lab at the Univ. of Nebraska. Mitochondrial, nuclear and chloroplast numbers, density and structure will be analyzed with tissue samples prepared as described by Segui-Samarro et al. (2008). Images will be collected using a 40x and a 60x objective, and Z-series images obtained by compiling 0.5 μm laser serial sections (for additional details see Cupp and Nielsen 2013).
We will examine plants at weekly time points and will determine organellar or nuclear localization of each replication protein in leaves, root tips, shoot apical meristems, and other tissues, to determine if localization is affected by tissue or stage of development. The effects of various stresses such as elevated temperature (>25°C), light (16 hr light/8 hr dark versus 12 hr light/12 hr dark and other ratios), reactive oxygen species, or photosynthesis or respiration inhibitors will also be tested to determine their effect on localization of the proteins.
Gene expression analysis. Very little information on coexpression is available in the ATTED Arabidopsis gene coexpression database (http://atted.jp) for any of the organelle replication protein genes except for PolIA as mentioned above using. Expression levels of each replication protein gene in different tissues and at different stages of growth will be analyzed using stably transformed plants expressing the native promoter-GUS constructs. GUS expression will be analyzed using published assays (Jefferson et al. 1987). In addition, transcript levels for each gene will be measured by quantitative reverse-transcriptase PCR (qRT-PCR; Cupp and Nielsen 2013). Total RNA will be purified from different plant tissues using a Qiagen RNA isolation kit, sampling once each week for 6 weeks after germination. DNA PolIA, PolIB, SSB1, and other replication genes will be analyzed to measure mRNA levels using a nuclear housekeeping gene such as actin 2 as control and primers that are specific for each organelle genome (Kumar and Bendich 2011; Cupp and Nielsen 2013).
We expect higher expression of the replication protein genes in actively growing cells (i.e. root tips and shoot apical meristem), but this may differ for each replication protein and may depend on if the plants are exposed to stress. We will develop comprehensive information on expression of these proteins under a range of growth and stress conditions and correlate with the localization analysis of each protein from above in order to determine the basis of the differences in growth and development between conditions. This will allow us to identify proteins that are coexpressed in similar tissues and organelles and provide a better understanding of proteins potentially involved in organelle genome replication during development.
Analysis of T-DNA insertion mutants. We have obtained several T-DNA insertion lines for the two DNA polymerase and SSB1 genes from ABRC (Cupp and Nielsen 2013). We have conducted backcrosses on the insertion lines and have confirmed that each has a single insertion. Initially our focus will be on SSB1 and the DNA PolIA mutants, followed by analysis of double PolIA/B mutants when they are confirmed. We are currently collecting data on the growth and development of PolIA mutant plants as we did for the PolIB insertion mutant (Cupp and Nielsen 2013). Our preliminary analysis indicates that the PolIA gene mutation is a simple recessive trait and heterozygous plants do not show an intermediate phenotype as was observed with the PolIB mutation (Cupp and Nielsen 2013). We have recently obtained seeds from homozygous PolIA/IB crosses, which will now be screened to obtain partial double mutants. We expect to obtain plants that are confirmed homozygous for insertion in one gene and heterozygous for the other gene insertion, in all possible combinations, using the same methods we have previously used (Cupp and Nielsen 2013). Plant growth will be monitored on a weekly basis and any differences as compared to wild-type plants recorded. Expression levels for each gene at weekly time points will be determined by qRT-PCR. Analysis of the crosses will allow determination of DNA polymerase gene expression levels and mtDNA and ctDNA copy numbers. This should provide important new insights into the effect of knocking down expression of both DNA polymerases.
RNAi to target the DNA polymerase genes. Because of the inability to obtain true double PolIA/IB mutants for the two DNA polymerase genes by us and others (Parent et al. 2011), we will also use an inducible RNAi approach to study the effect of decreased expression of each single gene and both in combination. We have obtained the pFGC1008 vector (Kerschen et al. 2004) for cloning short sequences specific to each organellar DNA polymerase gene. Each construct will be confirmed by DNA sequencing and transformed into plants by floral dipping as described above. Expression of each gene will be analyzed by quantitative reverse transcriptase PCR (qRT-PCR) to determine the degree of reduction in each transcript as described (Kerschen et al. 2004). Confirmed transformants for each DNA polymerase gene will then be crossed to obtain double RNAi plants where we will be able to study the effect of simultaneous downregulation of both genes by qRT-PCR. Alternatively, inducible RNAi constructs to target each DNA polymerase gene will be generated and analyzed as described (Burgos-Rivera et al. 2012). We do not expect to be able to completely knock down expression of both genes, but we do expect to see a dramatic effect in plants that show substantial reductions in both gene transcripts. We will measure the level of expression of each DNA polymerase gene and correlate the results with observations of growth rate, timing of shoot formation and flowering, and measurement of photosynthesis, respiration, and organelle DNA copy number as described below.
Determination of organelle DNA copy number by relative qPCR. The effect of the T-DNA insertions on mtDNA and ctDNA copy number will be determined using relative quantitative PCR (qPCR) with four separate primer pairs for distinct dispersed regions of the Arabidopsis mitochondrial genome and for the chloroplast genome (Preuten et al. 2010; Kumar and Bendich 2011; Cupp and Nielsen 2013). DNA will be purified from mutant and wild-type plants using a Qiagen DNA isolation kit. Relative levels of organelle DNA in comparison with a nuclearlocalized housekeeping gene control will be calculated (Cupp and Nielsen 2013). We found that the DNA PolIB mutant had a reduction in mtDNA levels but not in ctDNA compared to wild type plants (Cupp and Nielsen 2013). Since both DNA polymerases have been shown to be dual targeted to mitochondria and chloroplasts, we will look closely to determine whether the PolIA mutant exhibits a reduction in DNA in either organelle. We will also perform similar analysis of the double mutants. This will provide insight into any specific role that PolIA plays in DNA replication in each organelle. For example, if reduction in relative ctDNA levels is observed while mtDNA levels remain consistent in the PolIA mutant relative to wild type, this would suggest a prominent role for DNA PolIA in replication of the chloroplast genome. However, it is possible that the PolIA mutant will show either no reduction in DNA in either organelle if PolIB is primarily functional in the chloroplast, or a reduction in both mtDNA and ctDNA if PolIA is important for replication in both organelles. No alterations in mtDNA rearrangements, observed in some Arabidopsis organelle nucleoid protein mutants (Arrieta et al. 2009; Xu et al. 2011; 2012) have been observed in any of the mutants, so there does not appear to be a major shift in replication from one mechanism to another, such as a recombination-dependent mechanism. Different ages and tissues of seedlings grown under normal and various stress conditions such as drought, excess or restricted light, hydrogen peroxide or other ROS species, or growth inhibitors will be analyzed to allow determination of the factors that control expression of each protein.
Analysis of respiration and photosynthesis activity in wild type and mutant plants. To determine any differences in respiration and photosynthesis activity as related to changes in organelle genome copy number or other parameters, mutant and wild-type plants will be analyzed and compared (3-4 weeks post-germination). Respiration and photosynthesis measurements will be made following the methods we have recently used to characterize the DNA PolIB mutant (Cupp and Nielsen 2013). Briefly, seedlings will be germinated in glass scintillation vials containing 5 ml of growth medium. Gas exchange and carbon assimilation analysis will be conducted on the seedlings using a Li-Cor 6400XT Portable Photosynthesis System. Biological and technical replicates will be measured and statistical analysis will be performed to determine any significant differences between the mutant and wild-type plants.
Analysis of plants overexpressing the replication proteins. In addition to analysis of the TDNA insertion mutants that have no or low levels of the replication proteins, we will also analyze the effect of overexpression of the genes on plant growth, development, and organelle localization. This will be done using the constructs that have the CaMV35S promoter that drives expression of the genes. Analysis will be done as already described to measure respiration and photosynthesis and determine localization by confocal microscopy.
Summary, Expected Outcomes and Correlation with Future Work
Two major contributions will result from this research. First, this research will lead to confirmation of new examples of neofunctionalization in dual-targeted plant organelle proteins by identifying novel interactions with other organelle proteins in chloroplasts and mitochondria. Second, the results will contribute to a more complete picture of the organelle DNA replication machinery and provide details of expression of the proteins during plant development or as a result of stress. Production of stems, flowers and seeds is very energy-dependent, which puts demands on the organelles to produce energy and the building blocks required for development. A better understanding of the relationship between these demands and organelle DNA replication will be an added benefit of these studies.
The results from these studies will provide evidence for the degree of redundancy of the two DNA polymerases in replication of the mtDNA and ctDNA of Arabidopsis, and will provide clear data on when and in which plant organ each protein is expressed during plant development. The sequence analysis coupled with the protein interaction and nucleoid proteome analyses will provide evidence for neofunctionalization of organelle replication proteins. Any alterations in plant development will be correlated with changes in expression levels for each DNA polymerase gene in the single and double mutants. We expect that the partial double mutants will show a more pronounced phenotype than the single mutants, especially under stress conditions, and the results will help to understand more clearly the function of each DNA polymerase. We will develop a model for neofunctionalization of organelle replication proteins and their involvement in ctDNA and mtDNA maintenance.
We anticipate that new proteins that interact with the organelle replication proteins being studied will be identified. For example, if one of the RecA homologues interacts with SSB1 or one of the organellar DNA polymerases this may infer that these proteins are involved in recombinationdependent replication (RDR). Other proteins may be involved in regulation of ctDNA and mtDNA replication and control of genome copy number. Future studies will focus on the expression and function of these new proteins.
Anticipated Time Line
Year 1- Analyze gene expression and localization of proteins in stable transformed plants. Analyze sequences and initiate co-immunoprecipitation and proteome analysis experiments. Analyze the DNA PolIA and SSB1 mutants for photosynthesis and respiration and organelle DNA copy number. Begin screening of double DNA polymerase mutants. Year 2- Continue co-immunoprecipitation and characterization of interacting proteins, and nucleoid proteome analysis. Complete DNA copy number and gene expression analysis, protein localization, and screening of double DNA polymerase mutants. Year 3-Complete analysis of gene expression and interacting proteins in the organelle DNA replication complexes, analysis of mutant plants, prepare manuscripts and apply for new funding.
Educational Component
One Ph.D. student defended his dissertation on characterization of the Arabidopsis DNA PolIB gene last year (Cupp and Nielsen 2013). A fourth-year Ph.D. student has done some work on parts of the proposed project and on the Arabidopsis Twinkle DNA primase/DNA helicase gene (Diray-Arce et al. 2013) but is now working on a different project. A second-year Ph.D. student has conducted preliminary analysis of the DNA PolIA mutant and qPCR analysis of organelle genome copy numbers in the mutants, and will be supported by this grant. All three have trained undergraduate students in the techniques being using in our lab.
I am committed to the involvement of undergraduate students in research, and have mentored a large number of students (about 200) over the past 26 years. New students are paired with an experienced undergraduate or graduate student or work directly with me, and once they are trained they design and conduct their own experiments. I work closely with each student to make sure they understand the methods that they will use and how to interpret the data. Each student gives one formal report during our regular lab group meetings each semester. This involves providing background on their project and a summary of their results and conclusions, and discussion and input of ideas to help them with their project. The students interact often with other students and faculty in adjoining labs in the Life Sciences Building.
I often receive comments from former students that their experiences in my research laboratory prepared them well for graduate studies or employment. Several students have written to me that they found that they had much more experience with research techniques and were able to make contributions much more quickly on their new projects compared to their peers from other schools. A major benefit of working in my laboratory is that each student gains experience with a variety of methods including working with DNA, RNA and proteins. Recently graduated students who worked in my lab have gone on to Ph.D. programs at the Univ. of Pennsylvania, the Univ. of Colorado, Univ. of Virginia, and Scripps Institute among other places. Other former students are attending medical, pharmacy or dental school at various institutions, or are employed in biotechnology research or clinical laboratories.
All but one of the publications from our lab group in the last five years has a graduate student as first author, and all have at least one graduate student coauthor. One paper has an undergraduate student coauthor. Other publications over the past ten years have undergraduate coauthors, including one with an undergraduate student first author. It is expected that at least some of the publications resulting from this work will have undergraduate student coauthors, and most will have a graduate student coauthor, depending on their contributions to the experiments and the writing of the manuscripts. The postdoctoral researcher and graduate student should each have multiple first-author papers resulting from this work. Students will present their research at appropriate international meetings such as the annual meeting of the American Society for Biochemistry & Molecular Biology (ASBMB, San Diego, April 2016) or the American Society of Plant Biologists (ASPB, Minneapolis, July 2015 or Austin, July 2016) and at regional meetings.